| Molecular Weight: | 188.05 |
| Volume: | 192.251 |
| LogP: | 2.342 |
| LogD: | 2.107 |
| LogS: | -3.142 |
| # Rotatable Bonds: | 0 |
| TPSA: | 54.37 |
| # H-Bond Aceptor: | 3 |
| # H-Bond Donor: | 1 |
| # Rings: | 2 |
| # Heavy Atoms: | 3 |
| QED Drug-Likeness Score: | 0.674 |
| Synthetic Accessibility Score: | 2.267 |
| Fsp3: | 0.091 |
| Lipinski Rule-of-5: | Accepted |
| Pfizer Rule: | Accepted |
| GSK Rule: | Accepted |
| BMS Rule: | 0 |
| Golden Triangle Rule: | Rejected |
| Chelating Alert: | 0 |
| PAINS Alert: | 2 |
| Caco-2 Permeability: | -4.604 |
| MDCK Permeability: | 1.806890213629231e-05 |
| Pgp-inhibitor: | 0.017 |
| Pgp-substrate: | 0.0 |
| Human Intestinal Absorption (HIA): | 0.005 |
| 20% Bioavailability (F20%): | 0.003 |
| 30% Bioavailability (F30%): | 0.575 |
| Blood-Brain-Barrier Penetration (BBB): | 0.151 |
| Plasma Protein Binding (PPB): | 94.8353500366211% |
| Volume Distribution (VD): | 0.438 |
| Pgp-substrate: | 2.2197370529174805% |
| CYP1A2-inhibitor: | 0.975 |
| CYP1A2-substrate: | 0.541 |
| CYP2C19-inhibitor: | 0.583 |
| CYP2C19-substrate: | 0.079 |
| CYP2C9-inhibitor: | 0.666 |
| CYP2C9-substrate: | 0.669 |
| CYP2D6-inhibitor: | 0.885 |
| CYP2D6-substrate: | 0.235 |
| CYP3A4-inhibitor: | 0.716 |
| CYP3A4-substrate: | 0.16 |
| Clearance (CL): | 4.94 |
| Half-life (T1/2): | 0.594 |
| hERG Blockers: | 0.016 |
| Human Hepatotoxicity (H-HT): | 0.186 |
| Drug-inuced Liver Injury (DILI): | 0.465 |
| AMES Toxicity: | 0.908 |
| Rat Oral Acute Toxicity: | 0.951 |
| Maximum Recommended Daily Dose: | 0.899 |
| Skin Sensitization: | 0.89 |
| Carcinogencity: | 0.761 |
| Eye Corrosion: | 0.373 |
| Eye Irritation: | 0.991 |
| Respiratory Toxicity: | 0.471 |
 Natural Product: NPC285829
Natural Product: NPC285829| Natural Product ID: | NPC285829 |
| Common Name*: | Plumbagin |
| IUPAC Name: | 5-hydroxy-2-methylnaphthalene-1,4-dione |
| Synonyms: | plumbagin |
| Standard InCHIKey: | VCMMXZQDRFWYSE-UHFFFAOYSA-N |
| Standard InCHI: | InChI=1S/C11H8O3/c1-6-5-9(13)10-7(11(6)14)3-2-4-8(10)12/h2-5,12H,1H3 |
| SMILES: | CC1=CC(=O)c2c(cccc2O)C1=O |
| Synthetic Gene Cluster: | n.a. |
| ChEMBL Identifier: | CHEMBL295316 |
| PubChem CID: |
10205 |
| Chemical Classification**: |
|
*Note: the InCHIKey will be temporarily assigned as the "Common Name" if no IUPAC name or alternative short name is available.
**Note: the Chemical Classification was calculated by NPClassifier Version 1.5. Reference: PMID:34662515.
 Species Source
Species Source| Organism ID | Organism Name | Taxonomy Level | Family | SuperKingdom | Isolation Part | Collection Location | Collection Time | Reference |
|---|---|---|---|---|---|---|---|---|
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | stem | n.a. | DOI[10.1007/s004360050318] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | root | n.a. | DOI[10.1007/s004360050318] |
| NPO30483 | Plumbago rosea | Species | Plumbaginaceae | Eukaryota | n.a. | root | n.a. | DOI[10.2225/vol5-issue3-fulltext-4] |
| NPO25454 | Plumbago europaea | Species | Plumbaginaceae | Eukaryota | n.a. | root | n.a. | DOI[10.2225/vol5-issue3-fulltext-4] |
| NPO22192 | Cribraria cancellata | Species | Cribrariaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[14695806] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | leaf | n.a. |
PMID[14727919] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | root | n.a. |
PMID[14727919] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | leaf | n.a. |
PMID[14727919] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | root | n.a. |
PMID[14727919] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | xylem | n.a. |
PMID[14727919] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | root | n.a. |
PMID[14727919] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | bark | n.a. |
PMID[14727919] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | leaf | n.a. |
PMID[14727919] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | bark | n.a. |
PMID[14727919] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | xylem | n.a. |
PMID[14727919] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | bark | n.a. |
PMID[14727919] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | xylem | n.a. |
PMID[14727919] |
| NPO27122 | Plumbago scandens | Species | Plumbaginaceae | Eukaryota | n.a. | root | n.a. |
PMID[14762525] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | bark | Indonesia | n.a. |
PMID[15270571] |
| NPO13608 | Plumbago zeylanica | Species | Plumbaginaceae | Eukaryota | n.a. | root | n.a. |
PMID[16078700] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | pericarp | n.a. |
PMID[18496782] |
| NPO29119 | Nepenthes thorelii | Species | Nepenthaceae | Eukaryota | roots | n.a. | n.a. |
PMID[19299148] |
| NPO20721 | Halichondria panicea | Species | Halichondriidae | Eukaryota | n.a. | n.a. | n.a. |
PMID[20695474] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[22019229] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[22313254] |
| NPO33133 | barleria alluaudii | Species | Acanthaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[22313254] |
| NPO13608 | Plumbago zeylanica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[23848163] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[25466114] |
| NPO22069 | Persea major | Species | Lauraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[2614421] |
| NPO20721 | Halichondria panicea | Species | Halichondriidae | Eukaryota | n.a. | n.a. | n.a. |
PMID[27035556] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Flowers | n.a. | n.a. |
PMID[28737396] |
| NPO40718 | Indian beech tree | Strain | n.a. | n.a. | n.a. | n.a. | n.a. |
PMID[30108931] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | stem | n.a. |
PMID[9371362] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | root | n.a. |
PMID[9371362] |
| NPO29119 | Nepenthes thorelii | Species | Nepenthaceae | Eukaryota | n.a. | root | n.a. |
PMID[9581522] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | leaf | n.a. | Database[Article] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Essential Oil | n.a. | n.a. | Database[FooDB] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Fruit | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Fruit | n.a. | n.a. | Database[FooDB] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Leaf | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Leaf | n.a. | n.a. | Database[FooDB] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Pericarp | n.a. | n.a. | Database[FooDB] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Seed | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Seed | n.a. | n.a. | Database[FooDB] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | Seed | n.a. | n.a. | Database[FooDB] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | Shoot | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Stem Bark | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Testa | n.a. | n.a. | Database[FooDB] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | Embryo | n.a. | n.a. | Database[FooDB] |
| NPO2551 | Plumbagella micrantha | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO3770 | Drosera rotundifolia | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO22736.1 | Drosera peltata var. lunata | Varieties | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO14370 | Ceratostigma willmottianum | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO18595 | Ceratostigma plumbaginoides | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO13608 | Plumbago zeylanica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO31126 | Diospyros sp | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO18256 | Aconitum excelsum | Species | Ranunculaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO1775 | Euphorbia pekinensis | Species | Euphorbiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO31128 | Sparaxis sp | Species | Iridaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO25454 | Plumbago europaea | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO23045 | Lotus helleri | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO26780 | Dionaea muscipula | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO16662 | Plumbago indica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO26780 | Dionaea muscipula | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO3770 | Drosera rotundifolia | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO2551 | Plumbagella micrantha | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO16662 | Plumbago indica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO25454 | Plumbago europaea | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO1775 | Euphorbia pekinensis | Species | Euphorbiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO13608 | Plumbago zeylanica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO23045 | Lotus helleri | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO29133 | Sparaxis sp. | Species | Iridaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO18256 | Aconitum excelsum | Species | Ranunculaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO15529 | Diospyros sp. | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO18595 | Ceratostigma plumbaginoides | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO14370 | Ceratostigma willmottianum | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO22736.1 | Drosera peltata var. lunata | Varieties | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO1775 | Euphorbia pekinensis | Species | Euphorbiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO1775 | Euphorbia pekinensis | Species | Euphorbiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO22999 | Cussonia spicata | Species | Araliaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22831 | Isodon angustifolius | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO25454 | Plumbago europaea | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17858 | Juglans cinerea | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO20721 | Halichondria panicea | Species | Halichondriidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO19736 | Haplopappus glutinosus | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17623 | Melampodium linearilobum | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO21784 | Quillaja saponaria | Species | Quillajaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22069 | Persea major | Species | Lauraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO20094 | Loranthus micranthus | Species | Loranthaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17221 | Lagerstroemia parviflora | Species | Lythraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO27167 | Saguinus oedipus | Species | Cebidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18674 | Juglans regia | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO23517 | Heterotropa aspera | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO26647 | Ammannia baccifera | Species | Lythraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18256 | Aconitum excelsum | Species | Ranunculaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO13608 | Plumbago zeylanica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO2551 | Plumbagella micrantha | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18595 | Ceratostigma plumbaginoides | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO9170 | Pulsatilla pratensis | Species | Ranunculaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22192 | Cribraria cancellata | Species | Cribrariaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO23432 | Viguiera oaxacana | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO24158 | Phyllanthus anisolobus | Species | Leiothrichidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO26614 | Planchonella duclitan | Species | Sapotaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO6591 | Arthrospira maxima | Species | Microcoleaceae | Bacteria | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO29119 | Nepenthes thorelii | Species | Nepenthaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO6969 | Triphyophyllum peltatum | Species | Dioncophyllaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16657 | Diospyros maritima | Species | Ebenaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO25804 | Vernonia zanzibarensis | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO21383 | Cnicothamnus lorentzii | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO15106 | Agrostemma gracilis | Species | Caryophyllaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO29101 | Tanacetum larvatum | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO29276 | Juglans nigra | Species | Juglandaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17645.1 | Orobanche cernua var. cumana | Varieties | Orobanchaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18921 | Athrixia phylicoides | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO9038 | Aconitum burnatii | Species | Ranunculaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO15774 | Geranium pusillum | Species | Geraniaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18387 | Sedum formosanum | Species | Crassulaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO13717 | Aria elegans | Species | Rosaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO27122 | Plumbago scandens | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO1775 | Euphorbia pekinensis | Species | Euphorbiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO14370 | Ceratostigma willmottianum | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO27035 | Artemisia eriopoda | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO5057 | Ephestia kuehniella | Species | Pyralidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO3770 | Drosera rotundifolia | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18528 | Hyparrhenia hirta | Species | Poaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO26780 | Dionaea muscipula | Species | Droseraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO13113 | Ardisia kivuensis | Species | Primulaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO19238 | Choristoneura occidentalis | Species | Tortricidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22196 | Kickxia ramosissima | Species | Plantaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO23045 | Lotus helleri | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16662 | Plumbago indica | Species | Plumbaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO23865 | Scorzonera veratrifolia | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
☑ Note for Reference:
In addition to directly collecting NP source organism data from primary literature (where reference will provided as NCBI PMID or DOI links), NPASS also integrated them from below databases:
☉ UNPD: Universal Natural Products Database [PMID: 23638153].
☉ StreptomeDB: a database of streptomycetes natural products [PMID: 33051671].
☉ TM-MC: a database of medicinal materials and chemical compounds in Northeast Asian traditional medicine [PMID: 26156871].
☉ TCM@Taiwan: a Traditional Chinese Medicine database [PMID: 21253603].
☉ TCMID: a Traditional Chinese Medicine database [PMID: 29106634].
☉ TCMSP: The traditional Chinese medicine systems pharmacology database and analysis platform [PMID: 24735618].
☉ HerDing: a herb recommendation system to treat diseases using genes and chemicals [PMID: 26980517].
☉ MetaboLights: a metabolomics database [PMID: 27010336].
☉ FooDB: a database of constituents, chemistry and biology of food species [www.foodb.ca].
 NP Quantity Composition/Concentration
NP Quantity Composition/Concentration| Organism ID | NP ID | Organism Material Preparation | Organism Part | NP Quantity (Standard) | NP Quantity (Minimum) | NP Quantity (Maximum) | Quantity Unit | Reference |
|---|
☑ Note for Reference:
In addition to directly collecting NP quantitative data from primary literature (where reference will provided as NCBI PMID or DOI links), NPASS also integrated NP quantitative records for specific NP domains (e.g., NPS from foods or herbs) from domain-specific databases. These databases include:
☉ DUKE: Dr. Duke's Phytochemical and Ethnobotanical Databases.
☉ PHENOL EXPLORER: is the first comprehensive database on polyphenol content in foods [PMID: 24103452], its homepage can be accessed at here.
☉ FooDB: a database of constituents, chemistry and biology of food species [www.foodb.ca].
 Biological Activity
Biological Activity| Target ID | Target Type | Target Name | Target Organism | Activity Type | Activity Relation | Value | Unit | Reference |
|---|---|---|---|---|---|---|---|---|
| NPT143 | Individual Protein | DNA topoisomerase I | Homo sapiens | MIC | = | 50000.0 | nM | PMID[461002] |
| NPT91 | Cell Line | KB | Homo sapiens | ED50 | = | 0.3 | ug ml-1 | PMID[461008] |
| NPT1034 | Cell Line | Lu1 | Homo sapiens | ED50 | = | 0.3 | ug ml-1 | PMID[461008] |
| NPT858 | Cell Line | LNCaP | Homo sapiens | ED50 | = | 0.4 | ug ml-1 | PMID[461008] |
| NPT737 | Cell Line | HUVEC | Homo sapiens | ED50 | = | 0.4 | ug ml-1 | PMID[461008] |
| NPT4147 | Individual Protein | Mitogen-activated protein kinase kinase kinase 14 | Homo sapiens | IC50 | > | 100000.0 | nM | PMID[461012] |
| NPT4147 | Individual Protein | Mitogen-activated protein kinase kinase kinase 14 | Homo sapiens | Inhibition | = | 60.0 | % | PMID[461012] |
| NPT47 | Individual Protein | ATP-dependent DNA helicase Q1 | Homo sapiens | Potency | = | 22387.2 | nM | PMID[461014] |
| NPT149 | Individual Protein | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Homo sapiens | Potency | = | 12589.3 | nM | PMID[461014] |
| NPT149 | Individual Protein | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Homo sapiens | Potency | = | 39810.7 | nM | PMID[461013] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 8912.5 | nM | PMID[461014] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 6309.6 | nM | PMID[461013] |
| NPT1226 | Individual Protein | Caspase-7 | Homo sapiens | Potency | = | 19952.6 | nM | PMID[461014] |
| NPT50 | Individual Protein | Tyrosyl-DNA phosphodiesterase 1 | Homo sapiens | Potency | = | 39810.7 | nM | PMID[461013] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 2238.7 | nM | PMID[461013] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 891.3 | nM | PMID[461014] |
| NPT1424 | Individual Protein | Serine/threonine-protein kinase RAF | Homo sapiens | Potency | = | 25118.9 | nM | PMID[461014] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 562.3 | nM | PMID[461014] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 2511.9 | nM | PMID[461013] |
| NPT282 | Individual Protein | MAP kinase ERK2 | Homo sapiens | Potency | = | 7079.5 | nM | PMID[461014] |
| NPT531 | Individual Protein | Nuclear receptor ROR-gamma | Mus musculus | Potency | = | 12589.3 | nM | PMID[461013] |
| NPT531 | Individual Protein | Nuclear receptor ROR-gamma | Mus musculus | Potency | = | 5623.4 | nM | PMID[461014] |
| NPT53 | Individual Protein | 4'-phosphopantetheinyl transferase ffp | Bacillus subtilis | Potency | = | 35481.3 | nM | PMID[461013] |
| NPT583 | Individual Protein | Inositol monophosphatase 1 | Rattus norvegicus | Potency | = | 1995.3 | nM | PMID[461014] |
| NPT583 | Individual Protein | Inositol monophosphatase 1 | Rattus norvegicus | Potency | = | 25118.9 | nM | PMID[461013] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 631.0 | nM | PMID[461014] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 3162.3 | nM | PMID[461013] |
| NPT282 | Individual Protein | MAP kinase ERK2 | Homo sapiens | Potency | = | 19952.6 | nM | PMID[461013] |
| NPT282 | Individual Protein | MAP kinase ERK2 | Homo sapiens | Potency | = | 10000.0 | nM | PMID[461014] |
| NPT55 | Individual Protein | Putative fructose-1,6-bisphosphate aldolase | Giardia intestinalis | Potency | = | 111.9 | nM | PMID[461014] |
| NPT491 | Individual Protein | Dual specificity mitogen-activated protein kinase kinase 1 | Homo sapiens | Potency | = | 12589.3 | nM | PMID[461014] |
| NPT282 | Individual Protein | MAP kinase ERK2 | Homo sapiens | Potency | = | 5623.4 | nM | PMID[461014] |
| NPT531 | Individual Protein | Nuclear receptor ROR-gamma | Mus musculus | Potency | = | 2511.9 | nM | PMID[461014] |
| NPT151 | Individual Protein | 15-hydroxyprostaglandin dehydrogenase [NAD+] | Homo sapiens | Potency | = | 25118.9 | nM | PMID[461014] |
| NPT531 | Individual Protein | Nuclear receptor ROR-gamma | Mus musculus | Potency | = | 14125.4 | nM | PMID[461013] |
| NPT277 | Individual Protein | Caspase-1 | Homo sapiens | Potency | = | 31622.8 | nM | PMID[461013] |
| NPT539 | Individual Protein | Cellular tumor antigen p53 | Homo sapiens | Potency | = | 10000.0 | nM | PMID[461013] |
| NPT539 | Individual Protein | Cellular tumor antigen p53 | Homo sapiens | Potency | = | 3981.1 | nM | PMID[461014] |
| NPT539 | Individual Protein | Cellular tumor antigen p53 | Homo sapiens | Potency | = | 6309.6 | nM | PMID[461013] |
| NPT539 | Individual Protein | Cellular tumor antigen p53 | Homo sapiens | Potency | = | 794.3 | nM | PMID[461014] |
| NPT277 | Individual Protein | Caspase-1 | Homo sapiens | Potency | = | 12589.3 | nM | PMID[461014] |
| NPT58 | Individual Protein | Bloom syndrome protein | Homo sapiens | Potency | = | 31622.8 | nM | PMID[461014] |
| NPT109 | Individual Protein | Cytochrome P450 3A4 | Homo sapiens | Potency | = | 31622.8 | nM | PMID[461013] |
| NPT443 | Individual Protein | Histone acetyltransferase GCN5 | Homo sapiens | Potency | n.a. | 39810.7 | nM | PMID[461014] |
| NPT198 | Individual Protein | Vitamin D receptor | Homo sapiens | Potency | n.a. | 891.3 | nM | PMID[461014] |
| NPT198 | Individual Protein | Vitamin D receptor | Homo sapiens | Potency | n.a. | 7943.3 | nM | PMID[461013] |
| NPT199 | Individual Protein | DNA polymerase kappa | Homo sapiens | Potency | n.a. | 10000.0 | nM | PMID[461014] |
| NPT444 | Individual Protein | Ubiquitin carboxyl-terminal hydrolase 1 | Homo sapiens | Potency | n.a. | 17782.8 | nM | PMID[461014] |
| NPT444 | Individual Protein | Ubiquitin carboxyl-terminal hydrolase 1 | Homo sapiens | Potency | n.a. | 50118.7 | nM | PMID[461013] |
| NPT927 | Cell Line | PBMC | Homo sapiens | EC50 | = | 800.0 | nM | PMID[461015] |
| NPT367 | Cell Line | MDA-N | Homo sapiens | GI50 | n.a. | 1721.87 | nM | PMID[461016] |
| NPT368 | Cell Line | SN12C | Homo sapiens | GI50 | n.a. | 1940.89 | nM | PMID[461016] |
| NPT369 | Cell Line | ACHN | Homo sapiens | GI50 | n.a. | 1595.88 | nM | PMID[461016] |
| NPT370 | Cell Line | NCI-H23 | Homo sapiens | GI50 | n.a. | 1811.34 | nM | PMID[461016] |
| NPT371 | Cell Line | UO-31 | Homo sapiens | GI50 | n.a. | 1610.65 | nM | PMID[461016] |
| NPT116 | Cell Line | HL-60 | Homo sapiens | GI50 | n.a. | 152.41 | nM | PMID[461016] |
| NPT372 | Cell Line | HOP-92 | Homo sapiens | GI50 | n.a. | 464.52 | nM | PMID[461016] |
| NPT90 | Cell Line | DU-145 | Homo sapiens | GI50 | n.a. | 1648.16 | nM | PMID[461016] |
| NPT374 | Cell Line | SF-539 | Homo sapiens | GI50 | n.a. | 1914.26 | nM | PMID[461016] |
| NPT373 | Cell Line | SK-MEL-5 | Homo sapiens | GI50 | n.a. | 1621.81 | nM | PMID[461016] |
| NPT375 | Cell Line | Malme-3M | Homo sapiens | GI50 | n.a. | 966.05 | nM | PMID[461016] |
| NPT111 | Cell Line | K562 | Homo sapiens | GI50 | n.a. | 390.84 | nM | PMID[461016] |
| NPT376 | Cell Line | A498 | Homo sapiens | GI50 | n.a. | 5861.38 | nM | PMID[461016] |
| NPT112 | Cell Line | MOLT-4 | Homo sapiens | GI50 | n.a. | 235.5 | nM | PMID[461016] |
| NPT377 | Cell Line | OVCAR-3 | Homo sapiens | GI50 | n.a. | 1030.39 | nM | PMID[461016] |
| NPT379 | Cell Line | HOP-62 | Homo sapiens | GI50 | n.a. | 5520.77 | nM | PMID[461016] |
| NPT378 | Cell Line | NCI/ADR-RES | Homo sapiens | GI50 | n.a. | 1757.92 | nM | PMID[461016] |
| NPT381 | Cell Line | OVCAR-8 | Homo sapiens | GI50 | n.a. | 1690.44 | nM | PMID[461016] |
| NPT380 | Cell Line | U-251 | Homo sapiens | GI50 | n.a. | 6165.95 | nM | PMID[461016] |
| NPT382 | Cell Line | OVCAR-5 | Homo sapiens | GI50 | n.a. | 1940.89 | nM | PMID[461016] |
| NPT383 | Cell Line | SNB-19 | Homo sapiens | GI50 | n.a. | 12022.64 | nM | PMID[461016] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | GI50 | n.a. | 1918.67 | nM | PMID[461016] |
| NPT385 | Cell Line | SR | Homo sapiens | GI50 | n.a. | 1690.44 | nM | PMID[461016] |
| NPT384 | Cell Line | TK-10 | Homo sapiens | GI50 | n.a. | 1710.02 | nM | PMID[461016] |
| NPT323 | Cell Line | SW-620 | Homo sapiens | GI50 | n.a. | 1909.85 | nM | PMID[461016] |
| NPT455 | Cell Line | NCI-H522 | Homo sapiens | GI50 | n.a. | 1566.75 | nM | PMID[461016] |
| NPT386 | Cell Line | KM12 | Homo sapiens | GI50 | n.a. | 6165.95 | nM | PMID[461016] |
| NPT387 | Cell Line | M14 | Homo sapiens | GI50 | n.a. | 1862.09 | nM | PMID[461016] |
| NPT388 | Cell Line | NCI-H322M | Homo sapiens | GI50 | n.a. | 16292.96 | nM | PMID[461016] |
| NPT389 | Cell Line | RPMI-8226 | Homo sapiens | GI50 | n.a. | 891.25 | nM | PMID[461016] |
| NPT456 | Cell Line | OVCAR-4 | Homo sapiens | GI50 | n.a. | 1757.92 | nM | PMID[461016] |
| NPT390 | Cell Line | LOX IMVI | Homo sapiens | GI50 | n.a. | 1811.34 | nM | PMID[461016] |
| NPT457 | Cell Line | BT-549 | Homo sapiens | GI50 | n.a. | 1702.16 | nM | PMID[461016] |
| NPT147 | Cell Line | SK-MEL-2 | Homo sapiens | GI50 | n.a. | 1853.53 | nM | PMID[461016] |
| NPT391 | Cell Line | HCC 2998 | Homo sapiens | GI50 | n.a. | 2065.38 | nM | PMID[461016] |
| NPT81 | Cell Line | A549 | Homo sapiens | GI50 | n.a. | 11481.54 | nM | PMID[461016] |
| NPT392 | Cell Line | SNB-75 | Homo sapiens | GI50 | n.a. | 5508.08 | nM | PMID[461016] |
| NPT393 | Cell Line | HCT-116 | Homo sapiens | GI50 | n.a. | 1659.59 | nM | PMID[461016] |
| NPT148 | Cell Line | HCT-15 | Homo sapiens | GI50 | n.a. | 1702.16 | nM | PMID[461016] |
| NPT394 | Cell Line | EKVX | Homo sapiens | GI50 | n.a. | 1595.88 | nM | PMID[461016] |
| NPT395 | Cell Line | SF-268 | Homo sapiens | GI50 | n.a. | 2167.7 | nM | PMID[461016] |
| NPT306 | Cell Line | PC-3 | Homo sapiens | GI50 | n.a. | 2208.0 | nM | PMID[461016] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | GI50 | n.a. | 1790.61 | nM | PMID[461016] |
| NPT146 | Cell Line | SK-OV-3 | Homo sapiens | GI50 | n.a. | 14092.89 | nM | PMID[461016] |
| NPT396 | Cell Line | T47D | Homo sapiens | GI50 | n.a. | 433.51 | nM | PMID[461016] |
| NPT397 | Cell Line | NCI-H460 | Homo sapiens | GI50 | n.a. | 3639.15 | nM | PMID[461016] |
| NPT398 | Cell Line | UACC-62 | Homo sapiens | GI50 | n.a. | 1936.42 | nM | PMID[461016] |
| NPT308 | Cell Line | CAKI-1 | Homo sapiens | GI50 | n.a. | 1832.31 | nM | PMID[461016] |
| NPT400 | Cell Line | MDA-MB-435 | Homo sapiens | GI50 | n.a. | 1733.8 | nM | PMID[461016] |
| NPT458 | Cell Line | IGROV-1 | Homo sapiens | GI50 | n.a. | 1782.38 | nM | PMID[461016] |
| NPT399 | Cell Line | SF-295 | Homo sapiens | GI50 | n.a. | 11481.54 | nM | PMID[461016] |
| NPT401 | Cell Line | 786-0 | Homo sapiens | GI50 | n.a. | 1648.16 | nM | PMID[461016] |
| NPT402 | Cell Line | Hs-578T | Homo sapiens | GI50 | n.a. | 2910.72 | nM | PMID[461016] |
| NPT403 | Cell Line | UACC-257 | Homo sapiens | GI50 | n.a. | 1811.34 | nM | PMID[461016] |
| NPT404 | Cell Line | CCRF-CEM | Homo sapiens | GI50 | n.a. | 208.45 | nM | PMID[461016] |
| NPT139 | Cell Line | HT-29 | Homo sapiens | GI50 | n.a. | 1887.99 | nM | PMID[461016] |
| NPT405 | Cell Line | NCI-H226 | Homo sapiens | GI50 | n.a. | 6546.36 | nM | PMID[461016] |
| NPT170 | Cell Line | SK-MEL-28 | Homo sapiens | GI50 | n.a. | 1819.7 | nM | PMID[461016] |
| NPT407 | Cell Line | COLO 205 | Homo sapiens | GI50 | n.a. | 1862.09 | nM | PMID[461016] |
| NPT406 | Cell Line | RXF 393 | Homo sapiens | GI50 | n.a. | 1644.37 | nM | PMID[461016] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 5500.0 | nM | PMID[461017] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | IC50 | = | 3500.0 | nM | PMID[461017] |
| NPT784 | Cell Line | MDA-MB-468 | Homo sapiens | IC50 | = | 2500.0 | nM | PMID[461017] |
| NPT369 | Cell Line | ACHN | Homo sapiens | EC50 | = | 3700.0 | nM | PMID[461018] |
| NPT369 | Cell Line | ACHN | Homo sapiens | EC50 | = | 7300.0 | nM | PMID[461018] |
| NPT369 | Cell Line | ACHN | Homo sapiens | Ratio EC50 | = | 2.0 | n.a. | PMID[461018] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | IC50 | = | 3200.0 | nM | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | IC50 | = | 1800.0 | nM | PMID[461019] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | IC50 | = | 1800.0 | nM | PMID[461019] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | IC50 | = | 800.0 | nM | PMID[461019] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | Inhibition | = | 40.0 | % | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | Inhibition | = | 50.0 | % | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | Activity | = | 20.0 | % | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | Activity | = | 43.0 | % | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | Activity | = | 73.0 | % | PMID[461019] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | FC | = | 5.0 | n.a. | PMID[461019] |
| NPT134 | Cell Line | SK-BR-3 | Homo sapiens | FC | = | 4.0 | n.a. | PMID[461019] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | Activity | = | 70.0 | % | PMID[461019] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | Ratio IC50 | = | 2.0 | n.a. | PMID[461019] |
| NPT66 | Individual Protein | Acetylcholinesterase | Electrophorus electricus | Inhibition | = | 31.75 | % | PMID[461021] |
| NPT50 | Individual Protein | Tyrosyl-DNA phosphodiesterase 1 | Homo sapiens | Potency | n.a. | 2660.9 | nM | PMID[461013] |
| NPT50 | Individual Protein | Tyrosyl-DNA phosphodiesterase 1 | Homo sapiens | Potency | n.a. | 1158.2 | nM | PMID[461014] |
| NPT38 | Individual Protein | Signal transducer and activator of transcription 3 | Homo sapiens | Inhibition | = | 58.1 | % | PMID[461028] |
| NPT139 | Cell Line | HT-29 | Homo sapiens | IC50 | = | 4190.0 | nM | PMID[461028] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | IC50 | = | 21240.0 | nM | PMID[461028] |
| NPT90 | Cell Line | DU-145 | Homo sapiens | IC50 | = | 5230.0 | nM | PMID[461028] |
| NPT286 | Individual Protein | Receptor protein-tyrosine kinase erbB-2 | Homo sapiens | IC50 | = | 30800.0 | nM | PMID[461029] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 5100.0 | nM | PMID[461029] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 2840.0 | nM | PMID[461029] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 3350.0 | nM | PMID[461029] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | Activity | > | 90.0 | % | PMID[461029] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | n.a. | 28183.8 | nM | PMID[461014] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 12200.0 | nM | PMID[461030] |
| NPT165 | Cell Line | HeLa | Homo sapiens | IC50 | = | 59000.0 | nM | PMID[461030] |
| NPT2488 | Cell Line | MDA-MB-453 | Homo sapiens | IC50 | = | 34560.0 | nM | PMID[461030] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | IC50 | = | 51120.0 | nM | PMID[461030] |
| NPT306 | Cell Line | PC-3 | Homo sapiens | IC50 | = | 17190.0 | nM | PMID[461030] |
| NPT139 | Cell Line | HT-29 | Homo sapiens | IC50 | = | 26200.0 | nM | PMID[461030] |
| NPT660 | Cell Line | SW480 | Homo sapiens | IC50 | = | 12000.0 | nM | PMID[461032] |
| NPT660 | Cell Line | SW480 | Homo sapiens | IC50 | = | 7300.0 | nM | PMID[461032] |
| NPT323 | Cell Line | SW-620 | Homo sapiens | IC50 | = | 7700.0 | nM | PMID[461032] |
| NPT323 | Cell Line | SW-620 | Homo sapiens | IC50 | = | 7400.0 | nM | PMID[461032] |
| NPT393 | Cell Line | HCT-116 | Homo sapiens | IC50 | = | 20000.0 | nM | PMID[461032] |
| NPT323 | Cell Line | SW-620 | Homo sapiens | FC | = | 1.68 | n.a. | PMID[461032] |
| NPT1079 | Individual Protein | Histone acetyltransferase p300 | Homo sapiens | Kd | = | 2000.0 | nM | PMID[461033] |
| NPT1079 | Individual Protein | Histone acetyltransferase p300 | Homo sapiens | IC50 | = | 2000.0 | nM | PMID[461033] |
| NPT165 | Cell Line | HeLa | Homo sapiens | IC50 | = | 25500.0 | nM | PMID[461034] |
| NPT2393 | Cell Line | SAS | Homo sapiens | IC50 | = | 3000.0 | nM | PMID[461034] |
| NPT81 | Cell Line | A549 | Homo sapiens | IC50 | = | 3000.0 | nM | PMID[461036] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 3000.0 | nM | PMID[461036] |
| NPT170 | Cell Line | SK-MEL-28 | Homo sapiens | IC50 | = | 5000.0 | nM | PMID[461036] |
| NPT1079 | Individual Protein | Histone acetyltransferase p300 | Homo sapiens | IC50 | = | 20000.0 | nM | PMID[461037] |
| NPT1079 | Individual Protein | Histone acetyltransferase p300 | Homo sapiens | IC50 | = | 2000.0 | nM | PMID[461037] |
| NPT1080 | Individual Protein | Histone acetyltransferase PCAF | Homo sapiens | IC50 | = | 50000.0 | nM | PMID[461037] |
| NPT81 | Cell Line | A549 | Homo sapiens | MNCC | <= | 2.0 | uM | PMID[461039] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | GI | = | 32.8 | % | PMID[461040] |
| NPT65 | Cell Line | HepG2 | Homo sapiens | GI | = | 41.0 | % | PMID[461040] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | IC50 | = | 14230.0 | nM | PMID[461040] |
| NPT81 | Cell Line | A549 | Homo sapiens | IC50 | = | 12400.0 | nM | PMID[461040] |
| NPT65 | Cell Line | HepG2 | Homo sapiens | IC50 | = | 10660.0 | nM | PMID[461040] |
| NPT81 | Cell Line | A549 | Homo sapiens | FC | = | 1.34 | n.a. | PMID[461042] |
| NPT1527 | Cell Line | WRL68 | Homo sapiens | IC50 | = | 15360.0 | nM | PMID[461042] |
| NPT393 | Cell Line | HCT-116 | Homo sapiens | IC50 | = | 9800.0 | nM | PMID[461042] |
| NPT81 | Cell Line | A549 | Homo sapiens | IC50 | = | 8900.0 | nM | PMID[461042] |
| NPT65 | Cell Line | HepG2 | Homo sapiens | IC50 | = | 9170.0 | nM | PMID[461042] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | IC50 | = | 6500.0 | nM | PMID[461042] |
| NPT81 | Cell Line | A549 | Homo sapiens | GI | = | 60.7 | % | PMID[461042] |
| NPT65 | Cell Line | HepG2 | Homo sapiens | GI | = | 58.9 | % | PMID[461042] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | GI | = | 73.8 | % | PMID[461042] |
| NPT1374 | Cell Line | WI-38 | Homo sapiens | IC50 | = | 26710.0 | nM | PMID[461043] |
| NPT116 | Cell Line | HL-60 | Homo sapiens | IC50 | = | 3670.0 | nM | PMID[461043] |
| NPT116 | Cell Line | HL-60 | Homo sapiens | IC50 | = | 8250.0 | nM | PMID[461043] |
| NPT1603 | Individual Protein | Cytochrome P450 1A1 | Homo sapiens | Inhibition | = | 51.0 | % | PMID[461044] |
| NPT1604 | Individual Protein | Cytochrome P450 1B1 | Homo sapiens | Inhibition | = | 40.0 | % | PMID[461044] |
| NPT208 | Individual Protein | Cytochrome P450 1A2 | Homo sapiens | Inhibition | = | 55.0 | % | PMID[461044] |
| NPT110 | Individual Protein | Cytochrome P450 2D6 | Homo sapiens | Inhibition | = | 10.0 | % | PMID[461044] |
| NPT109 | Individual Protein | Cytochrome P450 3A4 | Homo sapiens | Inhibition | = | 15.0 | % | PMID[461044] |
| NPT212 | Individual Protein | Cytochrome P450 2C9 | Homo sapiens | Inhibition | = | 11.0 | % | PMID[461044] |
| NPT213 | Individual Protein | Cytochrome P450 2C19 | Homo sapiens | Inhibition | = | 12.0 | % | PMID[461044] |
| NPT1603 | Individual Protein | Cytochrome P450 1A1 | Homo sapiens | Inhibition | = | 86.0 | % | PMID[461044] |
| NPT1603 | Individual Protein | Cytochrome P450 1A1 | Homo sapiens | IC50 | = | 2770.0 | nM | PMID[461044] |
| NPT1603 | Individual Protein | Cytochrome P450 1A1 | Homo sapiens | Ki | = | 2560.0 | nM | PMID[461044] |
| NPT208 | Individual Protein | Cytochrome P450 1A2 | Homo sapiens | IC50 | = | 2460.0 | nM | PMID[461044] |
| NPT208 | Individual Protein | Cytochrome P450 1A2 | Homo sapiens | Ki | = | 2360.0 | nM | PMID[461044] |
| NPT1604 | Individual Protein | Cytochrome P450 1B1 | Homo sapiens | IC50 | = | 6150.0 | nM | PMID[461044] |
| NPT110 | Individual Protein | Cytochrome P450 2D6 | Homo sapiens | IC50 | > | 10000.0 | nM | PMID[461044] |
| NPT212 | Individual Protein | Cytochrome P450 2C9 | Homo sapiens | IC50 | > | 10000.0 | nM | PMID[461044] |
| NPT213 | Individual Protein | Cytochrome P450 2C19 | Homo sapiens | IC50 | > | 10000.0 | nM | PMID[461044] |
| NPT109 | Individual Protein | Cytochrome P450 3A4 | Homo sapiens | IC50 | > | 10000.0 | nM | PMID[461044] |
| NPT3441 | Individual Protein | Dual specificity mitogen-activated protein kinase kinase 7 | Homo sapiens | Kd | = | 14000.0 | nM | PMID[461045] |
| NPT3060 | Individual Protein | Dual specificity phosphatase Cdc25A | Homo sapiens | IC50 | = | 700.0 | nM | PMID[461045] |
| NPT1970 | Cell Line | THP-1 | Homo sapiens | IC50 | = | 1100.0 | nM | PMID[461045] |
| NPT2103 | Cell Line | MONO-MAC-6 | Homo sapiens | IC50 | = | 2600.0 | nM | PMID[461045] |
| NPT116 | Cell Line | HL-60 | Homo sapiens | IC50 | = | 1100.0 | nM | PMID[461045] |
| NPT2404 | Cell Line | SW982 | Homo sapiens | IC50 | = | 3600.0 | nM | PMID[461045] |
| NPT15 | Cell Line | Jurkat | Homo sapiens | IC50 | = | 950.0 | nM | PMID[461045] |
| NPT466 | Cell Line | U-937 | Homo sapiens | IC50 | = | 2400.0 | nM | PMID[461045] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 4100.0 | nM | PMID[461045] |
| NPT4246 | Cell Line | LS174T | Homo sapiens | IC50 | = | 9900.0 | nM | PMID[461045] |
| NPT81 | Cell Line | A549 | Homo sapiens | IC50 | = | 7200.0 | nM | PMID[461045] |
| NPT927 | Cell Line | PBMC | Homo sapiens | IC50 | = | 2700.0 | nM | PMID[461045] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | IC50 | = | 3500.0 | nM | PMID[461047] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | Activity | = | 20.0 | % | PMID[461047] |
| NPT784 | Cell Line | MDA-MB-468 | Homo sapiens | Activity | = | 25.0 | % | PMID[461047] |
| NPT784 | Cell Line | MDA-MB-468 | Homo sapiens | Activity | = | 35.0 | % | PMID[461047] |
| NPT784 | Cell Line | MDA-MB-468 | Homo sapiens | Activity | = | 20.0 | % | PMID[461047] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | Activity | = | 15.0 | % | PMID[461047] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | Activity | = | 10.0 | % | PMID[461047] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | CI | = | 0.9 | n.a. | PMID[461047] |
| NPT845 | Cell Line | BT-474 | Homo sapiens | CI | = | 0.6 | n.a. | PMID[461047] |
| NPT784 | Cell Line | MDA-MB-468 | Homo sapiens | CI | = | 0.5 | n.a. | PMID[461047] |
| NPT1079 | Individual Protein | Histone acetyltransferase p300 | Homo sapiens | IC50 | = | 20000.0 | nM | PMID[461048] |
| NPT1080 | Individual Protein | Histone acetyltransferase PCAF | Homo sapiens | IC50 | > | 50000.0 | nM | PMID[461048] |
| NPT467 | Individual Protein | Transient receptor potential cation channel subfamily A member 1 | Mus musculus | Activity | = | 50.0 | % | PMID[461049] |
| NPT346 | Individual Protein | Transient receptor potential cation channel subfamily A member 1 | Homo sapiens | Activity | = | 93.5 | pA/pF | PMID[461049] |
| NPT346 | Individual Protein | Transient receptor potential cation channel subfamily A member 1 | Homo sapiens | Activity | = | 65.0 | pA/pF | PMID[461049] |
| NPT346 | Individual Protein | Transient receptor potential cation channel subfamily A member 1 | Homo sapiens | EC50 | = | 460.0 | nM | PMID[461049] |
| NPT991 | Individual Protein | Trypanothione reductase | Trypanosoma cruzi | IC50 | = | 28000.0 | nM | PMID[461003] |
| NPT2 | Others | Unspecified | IC50 | = | 38000.0 | nM | PMID[461003] | |
| NPT2 | Others | Unspecified | IC50 | = | 2400.0 | nM | PMID[461004] | |
| NPT991 | Individual Protein | Trypanothione reductase | Trypanosoma cruzi | IC50 | = | 28000.0 | nM | PMID[461005] |
| NPT991 | Individual Protein | Trypanothione reductase | Trypanosoma cruzi | x fold | = | 525.0 | uM | PMID[461005] |
| NPT1018 | Organism | Trypanosoma brucei | Trypanosoma brucei | ED50 | = | 1.5 | uM | PMID[461005] |
| NPT991 | Individual Protein | Trypanothione reductase | Trypanosoma cruzi | Km | = | 15000.0 | nM | PMID[461005] |
| NPT991 | Individual Protein | Trypanothione reductase | Trypanosoma cruzi | Vmax | = | 0.31 | uM min-1mg-1 | PMID[461005] |
| NPT2 | Others | Unspecified | k cat/Km | = | 1.7 | 10e4 M-1 s-1 | PMID[461005] | |
| NPT2 | Others | Unspecified | Km | = | 40000.0 | nM | PMID[461005] | |
| NPT2 | Others | Unspecified | Vmax | = | 22.9 | uM min-1mg-1 | PMID[461005] | |
| NPT2 | Others | Unspecified | k cat/Km | = | 48.0 | 10e4 M-1 s-1 | PMID[461005] | |
| NPT2 | Others | Unspecified | Reduction | = | 26.1 | % | PMID[461005] | |
| NPT23850 | SINGLE PROTEIN | Dihydrolipoamide dehydrogenase | Sus scrofa | Km | = | 74000.0 | nM | PMID[461005] |
| NPT23850 | SINGLE PROTEIN | Dihydrolipoamide dehydrogenase | Sus scrofa | Vmax | = | 25.3 | uM min-1mg-1 | PMID[461005] |
| NPT23850 | SINGLE PROTEIN | Dihydrolipoamide dehydrogenase | Sus scrofa | k cat/Km | = | 29.0 | 10e4 M-1 s-1 | PMID[461005] |
| NPT23850 | SINGLE PROTEIN | Dihydrolipoamide dehydrogenase | Sus scrofa | Reduction | = | 15.4 | % | PMID[461005] |
| NPT27 | Others | Unspecified | Toxic dose | = | 3.0 | uM | PMID[461005] | |
| NPT27 | Others | Unspecified | Toxic dose | = | 10.0 | uM | PMID[461005] | |
| NPT1381 | Organism | Aedes aegypti | Aedes aegypti | Activity | = | 3.0 | ug ml-1 | PMID[461006] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | Ratio | >= | 4.0 | n.a. | PMID[461007] |
| NPT175 | Organism | Enterococcus faecalis | Enterococcus faecalis | MBC | = | 50.0 | ug ml-1 | PMID[461007] |
| NPT175 | Organism | Enterococcus faecalis | Enterococcus faecalis | MIC | = | 12.5 | ug.mL-1 | PMID[461007] |
| NPT1531 | Organism | Enterococcus faecium | Enterococcus faecium | MBC | = | 50.0 | ug ml-1 | PMID[461007] |
| NPT1531 | Organism | Enterococcus faecium | Enterococcus faecium | MIC | = | 12.5 | ug.mL-1 | PMID[461007] |
| NPT175 | Organism | Enterococcus faecalis | Enterococcus faecalis | MBC | = | 100.0 | ug ml-1 | PMID[461007] |
| NPT175 | Organism | Enterococcus faecalis | Enterococcus faecalis | MIC | = | 25.0 | ug.mL-1 | PMID[461007] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | MBC | = | 25.0 | ug ml-1 | PMID[461007] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | MIC | = | 3.13 | ug.mL-1 | PMID[461007] |
| NPT175 | Organism | Enterococcus faecalis | Enterococcus faecalis | Ratio | >= | 4.0 | n.a. | PMID[461007] |
| NPT1531 | Organism | Enterococcus faecium | Enterococcus faecium | Ratio | >= | 4.0 | n.a. | PMID[461007] |
| NPT729 | Organism | Micrococcus luteus | Micrococcus luteus | IC50 | = | 80.0 | ug.mL-1 | PMID[461008] |
| NPT729 | Organism | Micrococcus luteus | Micrococcus luteus | Activity | = | 600.0 | ug ml-1 | PMID[461008] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | IC50 | = | 100.0 | ug.mL-1 | PMID[461008] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | Activity | = | 600.0 | ug ml-1 | PMID[461008] |
| NPT602 | Organism | Mycobacterium smegmatis | Mycobacterium smegmatis | IC50 | = | 300.0 | ug.mL-1 | PMID[461008] |
| NPT602 | Organism | Mycobacterium smegmatis | Mycobacterium smegmatis | Activity | > | 1000.0 | ug ml-1 | PMID[461008] |
| NPT1513 | Organism | Mycobacterium avium | Mycobacterium avium | IC50 | = | 600.0 | ug.mL-1 | PMID[461008] |
| NPT1513 | Organism | Mycobacterium avium | Mycobacterium avium | Activity | > | 1000.0 | ug ml-1 | PMID[461008] |
| NPT20 | Organism | Candida albicans | Candida albicans | IC50 | = | 9.0 | ug.mL-1 | PMID[461008] |
| NPT85 | Organism | Filobasidiella neoformans | Cryptococcus neoformans | IC50 | = | 20.0 | ug.mL-1 | PMID[461008] |
| NPT85 | Organism | Filobasidiella neoformans | Cryptococcus neoformans | Activity | = | 40.0 | ug ml-1 | PMID[461008] |
| NPT312 | Organism | Saccharomyces cerevisiae | Saccharomyces cerevisiae | IC50 | = | 10.0 | ug.mL-1 | PMID[461008] |
| NPT312 | Organism | Saccharomyces cerevisiae | Saccharomyces cerevisiae | Activity | = | 40.0 | ug ml-1 | PMID[461008] |
| NPT21 | Organism | Aspergillus niger | Aspergillus niger | IC50 | = | 20.0 | ug.mL-1 | PMID[461008] |
| NPT21 | Organism | Aspergillus niger | Aspergillus niger | Activity | = | 400.0 | ug ml-1 | PMID[461008] |
| NPT20 | Organism | Candida albicans | Candida albicans | Activity | > | 1000.0 | ug ml-1 | PMID[461008] |
| NPT1381 | Organism | Aedes aegypti | Aedes aegypti | LC50 | = | 5.43 | ug.mL-1 | PMID[461010] |
| NPT1381 | Organism | Aedes aegypti | Aedes aegypti | LC90 | = | 6.56 | ug ml-1 | PMID[461010] |
| NPT1381 | Organism | Aedes aegypti | Aedes aegypti | Activity | = | 100.0 | % | PMID[461010] |
| NPT6 | Organism | Plasmodium falciparum | Plasmodium falciparum | IC50 | = | 270.0 | nM | PMID[461011] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 31622.8 | nM | PMID[461013] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 31622.8 | nM | PMID[461014] |
| NPT2 | Others | Unspecified | Potency | = | 2511.9 | nM | PMID[461014] | |
| NPT2 | Others | Unspecified | Potency | = | 5011.9 | nM | PMID[461013] | |
| NPT93 | Individual Protein | Survival motor neuron protein | Homo sapiens | Potency | = | 28183.8 | nM | PMID[461014] |
| NPT94 | Individual Protein | Aldehyde dehydrogenase 1A1 | Homo sapiens | Potency | = | 39810.7 | nM | PMID[461013] |
| NPT94 | Individual Protein | Aldehyde dehydrogenase 1A1 | Homo sapiens | Potency | = | 15848.9 | nM | PMID[461014] |
| NPT2 | Others | Unspecified | Potency | = | 15848.9 | nM | PMID[461014] | |
| NPT792 | Individual Protein | Arachidonate 15-lipoxygenase | Homo sapiens | Potency | = | 3162.3 | nM | PMID[461014] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 11220.2 | nM | PMID[461014] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 10000.0 | nM | PMID[461013] |
| NPT2 | Others | Unspecified | Potency | = | 28183.8 | nM | PMID[461014] | |
| NPT2 | Others | Unspecified | Potency | = | 14125.4 | nM | PMID[461013] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | EC50 | = | 800.0 | nM | PMID[461015] |
| NPT888 | Individual Protein | 78 kDa glucose-regulated protein | Homo sapiens | Potency | n.a. | 7943.3 | nM | PMID[461013] |
| NPT888 | Individual Protein | 78 kDa glucose-regulated protein | Homo sapiens | Potency | n.a. | 2511.9 | nM | PMID[461014] |
| NPT35 | Others | n.a. | LogP | = | 0.89 | n.a. | PMID[461017] | |
| NPT67 | Individual Protein | Cholinesterase | Equus caballus | Inhibition | = | 9.94 | % | PMID[461021] |
| NPT2747 | Organism | Achaea janata | Achaea janata | ED50 | = | 23.97 | microg/cm2 | PMID[461022] |
| NPT1175 | Organism | Spodoptera litura | Spodoptera litura | ED50 | = | 43.71 | microg/cm2 | PMID[461022] |
| NPT1365 | Organism | Trichoplusia ni | Trichoplusia ni | Activity | = | 12.7 | mg | PMID[461023] |
| NPT1365 | Organism | Trichoplusia ni | Trichoplusia ni | Activity | = | 3.5 | cm2 | PMID[461023] |
| NPT1365 | Organism | Trichoplusia ni | Trichoplusia ni | mortality | = | 7.0 | % | PMID[461023] |
| NPT1365 | Organism | Trichoplusia ni | Trichoplusia ni | DC50 | = | 3.3 | microg/cm2 | PMID[461023] |
| NPT1365 | Organism | Trichoplusia ni | Trichoplusia ni | FDI | = | 100.0 | % | PMID[461023] |
| NPT1499 | Organism | Erwinia amylovora | Erwinia amylovora | MIC | = | 100000.0 | nM | PMID[461024] |
| NPT2747 | Organism | Achaea janata | Achaea janata | Activity | = | 35.69 | % | PMID[461025] |
| NPT1175 | Organism | Spodoptera litura | Spodoptera litura | Activity | = | 26.0 | % | PMID[461025] |
| NPT2747 | Organism | Achaea janata | Achaea janata | mortality | < | 20.0 | % | PMID[461025] |
| NPT1175 | Organism | Spodoptera litura | Spodoptera litura | mortality | < | 20.0 | % | PMID[461025] |
| NPT2747 | Organism | Achaea janata | Achaea janata | LC50 | = | 252.52 | microg/cm2 | PMID[461025] |
| NPT1175 | Organism | Spodoptera litura | Spodoptera litura | LC50 | = | 335.52 | microg/cm2 | PMID[461025] |
| NPT790 | Organism | Mycobacterium tuberculosis H37Rv | Mycobacterium tuberculosis H37Rv | MIC | = | 42.3 | ug.mL-1 | PMID[461026] |
| NPT790 | Organism | Mycobacterium tuberculosis H37Rv | Mycobacterium tuberculosis H37Rv | MIC | = | 40.3 | ug.mL-1 | PMID[461026] |
| NPT32 | Organism | Mus musculus | Mus musculus | FC | > | 3.0 | n.a. | PMID[461027] |
| NPT32 | Organism | Mus musculus | Mus musculus | Activity | = | 77.0 | % | PMID[461027] |
| NPT32 | Organism | Mus musculus | Mus musculus | Activity | = | 0.3 | % | PMID[461027] |
| NPT29 | Organism | Rattus norvegicus | Rattus norvegicus | F | = | 38.0 | % | PMID[461027] |
| NPT2 | Others | Unspecified | Potency | n.a. | 3162.3 | nM | PMID[461013] | |
| NPT2 | Others | Unspecified | Potency | n.a. | 6309.6 | nM | PMID[461014] | |
| NPT2 | Others | Unspecified | Potency | n.a. | 3981.1 | nM | PMID[461013] | |
| NPT2 | Others | Unspecified | Potency | n.a. | 3548.1 | nM | PMID[461014] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | IC50 | = | 43400.0 | nM | PMID[461030] |
| NPT2 | Others | Unspecified | Inhibition | = | 50.0 | % | PMID[461032] | |
| NPT2 | Others | Unspecified | Inhibition | = | 33.0 | % | PMID[461032] | |
| NPT2 | Others | Unspecified | Inhibition | = | 20.0 | % | PMID[461032] | |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | IZ | = | 12.0 | mm | PMID[461034] |
| NPT20 | Organism | Candida albicans | Candida albicans | IZ | = | 12.0 | mm | PMID[461034] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | MIC | = | 125.0 | ug.mL-1 | PMID[461034] |
| NPT20 | Organism | Candida albicans | Candida albicans | MIC | = | 125.0 | ug.mL-1 | PMID[461034] |
| NPT88 | Organism | Mycobacterium tuberculosis | Mycobacterium tuberculosis | MIC | = | 4.0 | ug.mL-1 | PMID[461035] |
| NPT88 | Organism | Mycobacterium tuberculosis | Mycobacterium tuberculosis | MIC | = | 32.0 | ug.mL-1 | PMID[461035] |
| NPT88 | Organism | Mycobacterium tuberculosis | Mycobacterium tuberculosis | MIC | = | 128.0 | ug.mL-1 | PMID[461035] |
| NPT2 | Others | Unspecified | Potency | n.a. | 8912.5 | nM | PMID[461014] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | IC50 | = | 4000.0 | nM | PMID[461036] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | IC50 | = | 1000.0 | nM | PMID[461036] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | IC50 | = | 5000.0 | nM | PMID[461036] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | MIC | = | 4.0 | ug.mL-1 | PMID[461038] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | MIC | = | 2.0 | ug.mL-1 | PMID[461038] |
| NPT2 | Others | Unspecified | Inhibition | = | 58.0 | % | PMID[461038] | |
| NPT2 | Others | Unspecified | Inhibition | = | 45.0 | % | PMID[461038] | |
| NPT2 | Others | Unspecified | Inhibition | = | 33.0 | % | PMID[461038] | |
| NPT2 | Others | Unspecified | Inhibition | = | 26.0 | % | PMID[461038] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | MIC | = | 32.0 | ug.mL-1 | PMID[461038] |
| NPT2 | Others | Unspecified | Inhibition | = | 52.0 | % | PMID[461039] | |
| NPT2 | Others | Unspecified | Inhibition | = | 74.0 | % | PMID[461039] | |
| NPT2 | Others | Unspecified | Inhibition | = | 38.0 | % | PMID[461039] | |
| NPT2 | Others | Unspecified | Inhibition | = | 67.0 | % | PMID[461039] | |
| NPT2 | Others | Unspecified | Ratio IC50 | = | 0.9 | n.a. | PMID[461040] | |
| NPT2 | Others | Unspecified | Ratio IC50 | = | 0.7 | n.a. | PMID[461040] | |
| NPT35 | Others | n.a. | Drug degradation | > | 95.0 | % | PMID[461040] | |
| NPT580 | Organism | Trypanosoma cruzi | Trypanosoma cruzi | IC50 | = | 1380.0 | nM | PMID[461041] |
| NPT23852 | CELL-LINE | HK-2 | Homo sapiens | IC50 | = | 23580.0 | nM | PMID[461042] |
| NPT2 | Others | Unspecified | Ratio IC50 | = | 7.28 | n.a. | PMID[461043] | |
| NPT911 | Individual Protein | Dual specificity phosphatase Cdc25B | Homo sapiens | IC50 | = | 3100.0 | nM | PMID[461045] |
| NPT21773 | SINGLE PROTEIN | Thiosulfate sulfurtransferase | Homo sapiens | IC50 | = | 60.0 | nM | PMID[461046] |
| NPT2 | Others | Unspecified | IC50 | > | 63000.0 | nM | PMID[461046] | |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | IC50 | = | 290.0 | nM | PMID[461046] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | IC50 | = | 3800.0 | nM | PMID[461046] |
| NPT21774 | PROTEIN COMPLEX | HSP60/HSP10 | Homo sapiens | IC50 | = | 6300.0 | nM | PMID[461046] |
| NPT21775 | SINGLE PROTEIN | 60 kDa chaperonin | Escherichia coli (strain K12) | IC50 | > | 250000.0 | nM | PMID[461046] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | Inhibition | = | 87.0 | % | PMID[461046] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | Inhibition | = | 92.0 | % | PMID[461046] |
| NPT23853 | CELL-LINE | MCF7/PTX | Homo sapiens | Activity | = | 20.0 | % | PMID[461047] |
| NPT23853 | CELL-LINE | MCF7/PTX | Homo sapiens | IC50 | = | 3500.0 | nM | PMID[461047] |
| NPT35 | Others | n.a. | LogP | = | 3.69 | n.a. | PMID[461049] | |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | IC50 | > | 20000.0 | nM | PMID[461050] |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | IC50 | > | 19952.62 | nM | PMID[461050] |
☑ Note for Activity Records:
☉ The quantitative biological activities were primarily integrated from ChEMBL (Version-30) database and were also directly collected from PubMed literature. PubMed PMID was provided as the reference link for each activity record.
 Chemically structural similarity: I. Similar Active Natural Products in NPASS
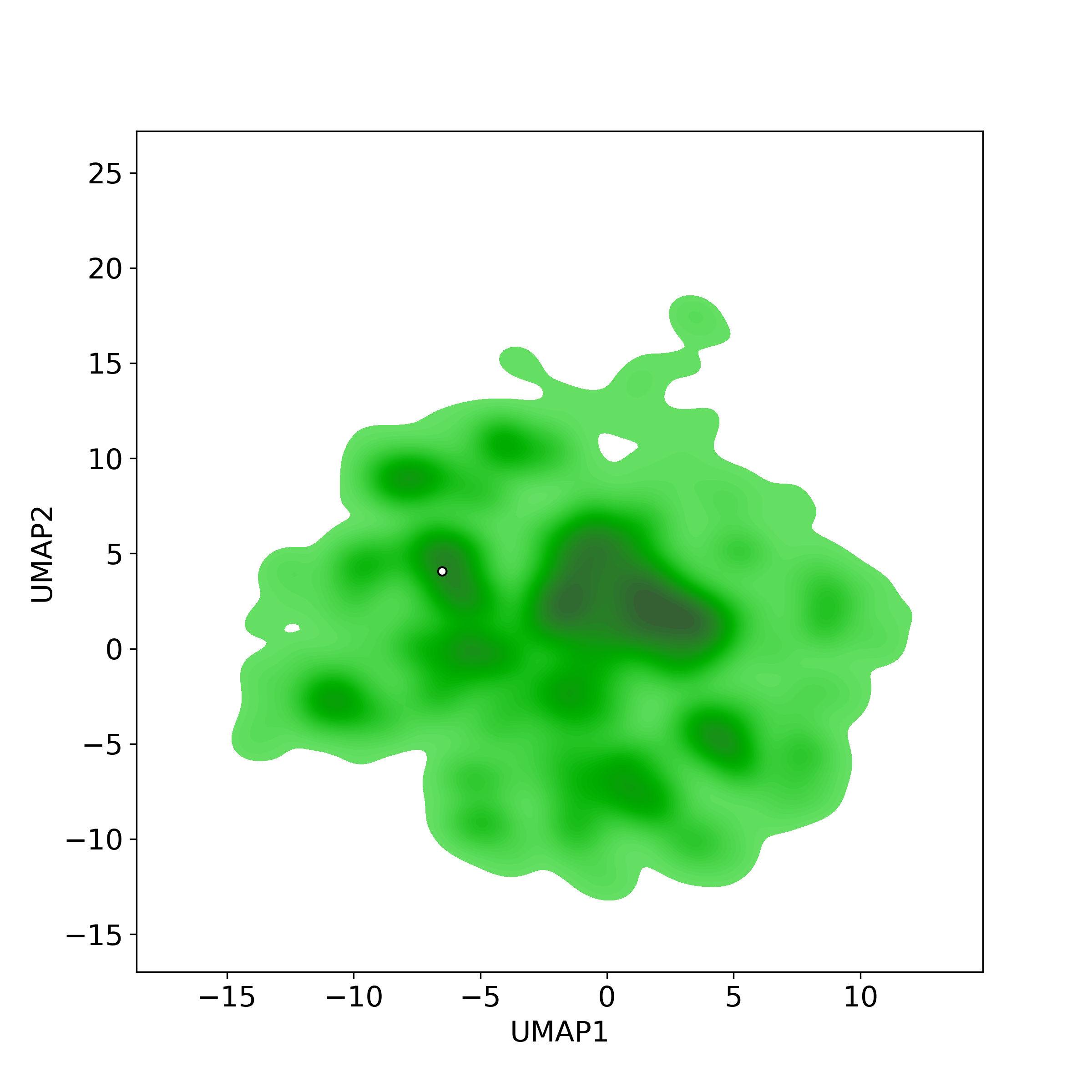
Chemically structural similarity: I. Similar Active Natural Products in NPASSTop-200 similar NPs were calculated against the active-NP-set (includes 4,3285 NPs with experimentally-derived bioactivity available in NPASS)
Similarity level is defined by Tanimoto coefficient (Tc) between two molecules. Tc lies between [0, 1] where '1' indicates the highest similarity. What is Tanimoto coefficient 
● The left chart: Distribution of similarity level between NPC285829 and all remaining natural products in the NPASS database.
● The right table: Most similar natural products (Tc>=0.56 or Top200).
| Similarity Score | Similarity Level | Natural Product ID |
|---|---|---|
| 1.0 | High Similarity | NPC206778 |
| 0.9739 | High Similarity | NPC307174 |
| 0.9643 | High Similarity | NPC108288 |
| 0.9573 | High Similarity | NPC310540 |
| 0.9412 | High Similarity | NPC306765 |
| 0.9412 | High Similarity | NPC3224 |
| 0.9412 | High Similarity | NPC231774 |
| 0.9402 | High Similarity | NPC300274 |
| 0.9322 | High Similarity | NPC375356 |
| 0.9256 | High Similarity | NPC237225 |
| 0.9244 | High Similarity | NPC142956 |
| 0.9224 | High Similarity | NPC232178 |
| 0.9211 | High Similarity | NPC161304 |
| 0.918 | High Similarity | NPC34414 |
| 0.918 | High Similarity | NPC146647 |
| 0.9167 | High Similarity | NPC103540 |
| 0.913 | High Similarity | NPC477453 |
| 0.9106 | High Similarity | NPC117609 |
| 0.9098 | High Similarity | NPC282923 |
| 0.9091 | High Similarity | NPC96915 |
| 0.9091 | High Similarity | NPC1991 |
| 0.9091 | High Similarity | NPC51037 |
| 0.9083 | High Similarity | NPC234890 |
| 0.9083 | High Similarity | NPC74507 |
| 0.9043 | High Similarity | NPC240163 |
| 0.9032 | High Similarity | NPC244699 |
| 0.9024 | High Similarity | NPC99731 |
| 0.8983 | High Similarity | NPC216297 |
| 0.8983 | High Similarity | NPC7151 |
| 0.8983 | High Similarity | NPC473662 |
| 0.896 | High Similarity | NPC110609 |
| 0.896 | High Similarity | NPC242358 |
| 0.896 | High Similarity | NPC246693 |
| 0.8952 | High Similarity | NPC198305 |
| 0.8952 | High Similarity | NPC55949 |
| 0.8943 | High Similarity | NPC165257 |
| 0.8943 | High Similarity | NPC48248 |
| 0.8929 | High Similarity | NPC224584 |
| 0.8926 | High Similarity | NPC68756 |
| 0.8926 | High Similarity | NPC173978 |
| 0.8926 | High Similarity | NPC152525 |
| 0.8889 | High Similarity | NPC125252 |
| 0.8889 | High Similarity | NPC269414 |
| 0.8889 | High Similarity | NPC53896 |
| 0.8889 | High Similarity | NPC171968 |
| 0.888 | High Similarity | NPC205992 |
| 0.8871 | High Similarity | NPC278928 |
| 0.8862 | High Similarity | NPC471530 |
| 0.8843 | High Similarity | NPC182646 |
| 0.8839 | High Similarity | NPC34715 |
| 0.8819 | High Similarity | NPC50924 |
| 0.8814 | High Similarity | NPC41567 |
| 0.881 | High Similarity | NPC309430 |
| 0.881 | High Similarity | NPC58685 |
| 0.88 | High Similarity | NPC136588 |
| 0.88 | High Similarity | NPC475741 |
| 0.88 | High Similarity | NPC199253 |
| 0.878 | High Similarity | NPC160499 |
| 0.876 | High Similarity | NPC282577 |
| 0.875 | High Similarity | NPC281513 |
| 0.875 | High Similarity | NPC283088 |
| 0.875 | High Similarity | NPC22222 |
| 0.874 | High Similarity | NPC115458 |
| 0.874 | High Similarity | NPC225051 |
| 0.873 | High Similarity | NPC205360 |
| 0.8689 | High Similarity | NPC135062 |
| 0.8682 | High Similarity | NPC193358 |
| 0.8678 | High Similarity | NPC120545 |
| 0.8661 | High Similarity | NPC70622 |
| 0.8651 | High Similarity | NPC31799 |
| 0.8629 | High Similarity | NPC136342 |
| 0.8629 | High Similarity | NPC227741 |
| 0.8629 | High Similarity | NPC295202 |
| 0.8629 | High Similarity | NPC49647 |
| 0.8621 | High Similarity | NPC231717 |
| 0.8618 | High Similarity | NPC236189 |
| 0.8615 | High Similarity | NPC238108 |
| 0.8607 | High Similarity | NPC217756 |
| 0.8607 | High Similarity | NPC91478 |
| 0.8605 | High Similarity | NPC80035 |
| 0.8605 | High Similarity | NPC161632 |
| 0.8595 | High Similarity | NPC178395 |
| 0.8595 | High Similarity | NPC159760 |
| 0.8595 | High Similarity | NPC272454 |
| 0.8595 | High Similarity | NPC26433 |
| 0.8595 | High Similarity | NPC35856 |
| 0.8595 | High Similarity | NPC115188 |
| 0.8595 | High Similarity | NPC190971 |
| 0.8595 | High Similarity | NPC301987 |
| 0.8595 | High Similarity | NPC292665 |
| 0.8595 | High Similarity | NPC244994 |
| 0.8595 | High Similarity | NPC222876 |
| 0.8595 | High Similarity | NPC179092 |
| 0.8594 | High Similarity | NPC294226 |
| 0.8594 | High Similarity | NPC52407 |
| 0.8594 | High Similarity | NPC254603 |
| 0.8583 | High Similarity | NPC233165 |
| 0.8583 | High Similarity | NPC184579 |
| 0.8583 | High Similarity | NPC17083 |
| 0.856 | High Similarity | NPC45537 |
| 0.855 | High Similarity | NPC173980 |
| 0.855 | High Similarity | NPC61590 |
| 0.8548 | High Similarity | NPC123506 |
| 0.8548 | High Similarity | NPC131799 |
| 0.8538 | High Similarity | NPC272268 |
| 0.8538 | High Similarity | NPC315275 |
| 0.8538 | High Similarity | NPC155211 |
| 0.8538 | High Similarity | NPC141934 |
| 0.8538 | High Similarity | NPC474813 |
| 0.8537 | High Similarity | NPC275145 |
| 0.8537 | High Similarity | NPC477153 |
| 0.8527 | High Similarity | NPC44437 |
| 0.8527 | High Similarity | NPC288089 |
| 0.8525 | High Similarity | NPC198336 |
| 0.8516 | High Similarity | NPC477406 |
| 0.8509 | High Similarity | NPC141523 |
| 0.8504 | High Similarity | NPC31539 |
| 0.8492 | Intermediate Similarity | NPC179898 |
| 0.8492 | Intermediate Similarity | NPC96024 |
| 0.8485 | Intermediate Similarity | NPC299405 |
| 0.8485 | Intermediate Similarity | NPC306835 |
| 0.8485 | Intermediate Similarity | NPC476477 |
| 0.8485 | Intermediate Similarity | NPC29771 |
| 0.8485 | Intermediate Similarity | NPC216312 |
| 0.8485 | Intermediate Similarity | NPC295339 |
| 0.8485 | Intermediate Similarity | NPC471602 |
| 0.8485 | Intermediate Similarity | NPC256463 |
| 0.8485 | Intermediate Similarity | NPC111422 |
| 0.848 | Intermediate Similarity | NPC15837 |
| 0.848 | Intermediate Similarity | NPC254492 |
| 0.8473 | Intermediate Similarity | NPC305060 |
| 0.8473 | Intermediate Similarity | NPC53414 |
| 0.8473 | Intermediate Similarity | NPC53206 |
| 0.8473 | Intermediate Similarity | NPC245923 |
| 0.8468 | Intermediate Similarity | NPC199273 |
| 0.8468 | Intermediate Similarity | NPC265910 |
| 0.8468 | Intermediate Similarity | NPC91475 |
| 0.8462 | Intermediate Similarity | NPC258502 |
| 0.8455 | Intermediate Similarity | NPC283514 |
| 0.8455 | Intermediate Similarity | NPC244351 |
| 0.8455 | Intermediate Similarity | NPC46634 |
| 0.8443 | Intermediate Similarity | NPC472046 |
| 0.8438 | Intermediate Similarity | NPC72669 |
| 0.8438 | Intermediate Similarity | NPC474517 |
| 0.8425 | Intermediate Similarity | NPC25736 |
| 0.8421 | Intermediate Similarity | NPC32749 |
| 0.8421 | Intermediate Similarity | NPC327916 |
| 0.8421 | Intermediate Similarity | NPC241349 |
| 0.8421 | Intermediate Similarity | NPC42262 |
| 0.8421 | Intermediate Similarity | NPC220496 |
| 0.8421 | Intermediate Similarity | NPC147542 |
| 0.8421 | Intermediate Similarity | NPC37992 |
| 0.8417 | Intermediate Similarity | NPC160199 |
| 0.8413 | Intermediate Similarity | NPC72667 |
| 0.8413 | Intermediate Similarity | NPC470841 |
| 0.8409 | Intermediate Similarity | NPC416 |
| 0.8409 | Intermediate Similarity | NPC13715 |
| 0.8409 | Intermediate Similarity | NPC4214 |
| 0.8409 | Intermediate Similarity | NPC61398 |
| 0.8397 | Intermediate Similarity | NPC254847 |
| 0.8397 | Intermediate Similarity | NPC86524 |
| 0.8385 | Intermediate Similarity | NPC282780 |
| 0.8385 | Intermediate Similarity | NPC166480 |
| 0.8358 | Intermediate Similarity | NPC471444 |
| 0.8358 | Intermediate Similarity | NPC84273 |
| 0.8358 | Intermediate Similarity | NPC257003 |
| 0.8346 | Intermediate Similarity | NPC242994 |
| 0.8346 | Intermediate Similarity | NPC767 |
| 0.8346 | Intermediate Similarity | NPC477408 |
| 0.8346 | Intermediate Similarity | NPC95537 |
| 0.8346 | Intermediate Similarity | NPC26924 |
| 0.8346 | Intermediate Similarity | NPC114620 |
| 0.8346 | Intermediate Similarity | NPC103337 |
| 0.8346 | Intermediate Similarity | NPC138099 |
| 0.8346 | Intermediate Similarity | NPC1268 |
| 0.8346 | Intermediate Similarity | NPC247250 |
| 0.8333 | Intermediate Similarity | NPC471452 |
| 0.8333 | Intermediate Similarity | NPC471905 |
| 0.8333 | Intermediate Similarity | NPC71610 |
| 0.8319 | Intermediate Similarity | NPC118288 |
| 0.8319 | Intermediate Similarity | NPC163154 |
| 0.8319 | Intermediate Similarity | NPC276111 |
| 0.8308 | Intermediate Similarity | NPC314048 |
| 0.8306 | Intermediate Similarity | NPC164947 |
| 0.8296 | Intermediate Similarity | NPC474300 |
| 0.8295 | Intermediate Similarity | NPC85342 |
| 0.8295 | Intermediate Similarity | NPC93015 |
| 0.8293 | Intermediate Similarity | NPC165197 |
| 0.8284 | Intermediate Similarity | NPC315578 |
| 0.8284 | Intermediate Similarity | NPC156872 |
| 0.8284 | Intermediate Similarity | NPC169452 |
| 0.8284 | Intermediate Similarity | NPC181560 |
| 0.8281 | Intermediate Similarity | NPC171460 |
| 0.8281 | Intermediate Similarity | NPC92624 |
| 0.8268 | Intermediate Similarity | NPC84672 |
| 0.8268 | Intermediate Similarity | NPC69424 |
| 0.8268 | Intermediate Similarity | NPC473787 |
| 0.8268 | Intermediate Similarity | NPC176130 |
| 0.8268 | Intermediate Similarity | NPC78364 |
| 0.8268 | Intermediate Similarity | NPC181715 |
 Chemically structural similarity: II. Similar Clinical/Approved Drugs
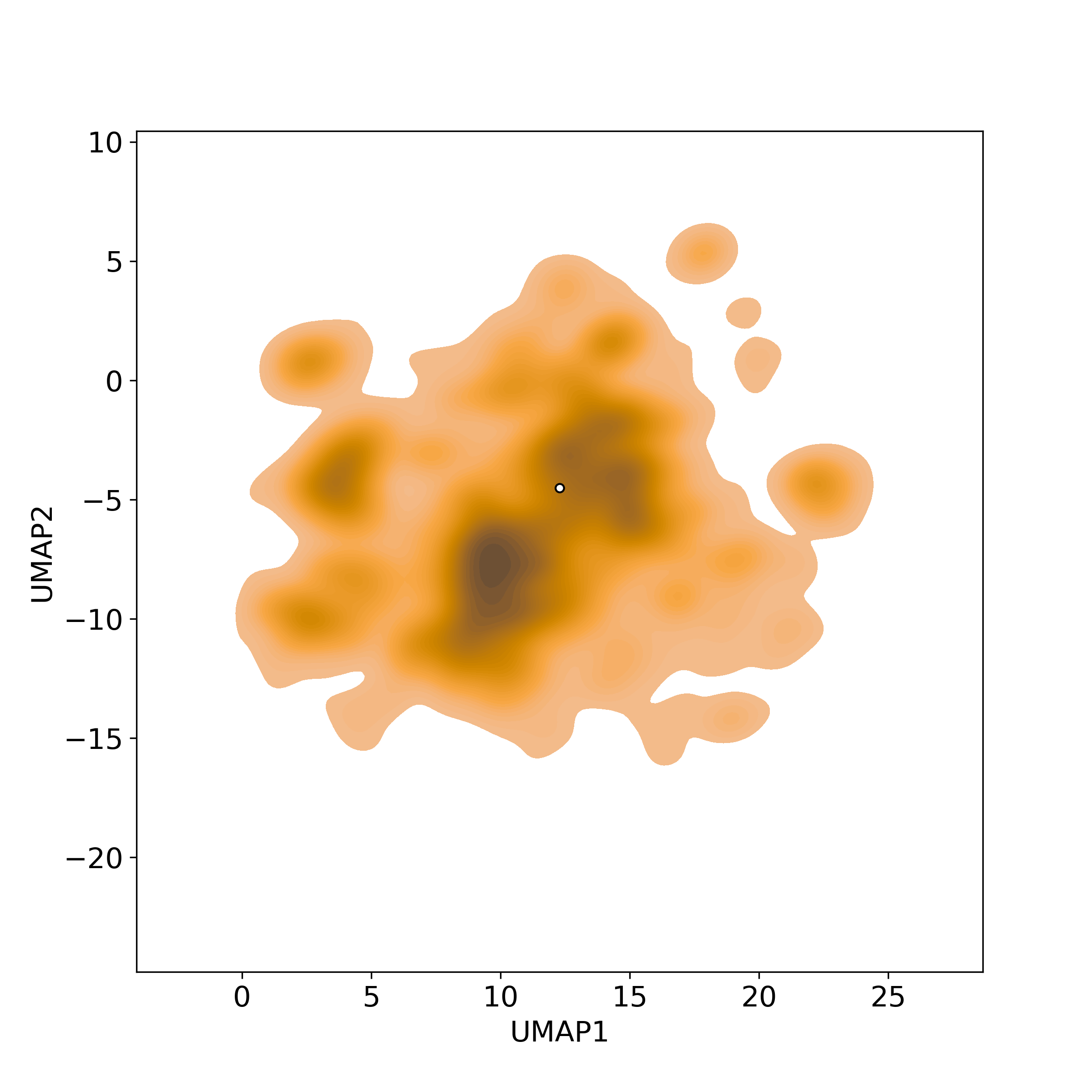
Chemically structural similarity: II. Similar Clinical/Approved DrugsSimilarity level is defined by Tanimoto coefficient (Tc) between two molecules.
● The left chart: Distribution of similarity level between NPC285829 and all drugs/candidates.
● The right table: Most similar clinical/approved drugs (Tc>=0.56 or Top200).
| Similarity Score | Similarity Level | Drug ID | Developmental Stage |
|---|---|---|---|
| 0.9643 | High Similarity | NPD405 | Clinical (unspecified phase) |
| 0.9167 | High Similarity | NPD1470 | Approved |
| 0.8992 | High Similarity | NPD1201 | Approved |
| 0.8397 | Intermediate Similarity | NPD1509 | Clinical (unspecified phase) |
| 0.812 | Intermediate Similarity | NPD9266 | Approved |
| 0.812 | Intermediate Similarity | NPD74 | Approved |
| 0.8034 | Intermediate Similarity | NPD9267 | Approved |
| 0.8034 | Intermediate Similarity | NPD9264 | Approved |
| 0.8034 | Intermediate Similarity | NPD9263 | Approved |
| 0.8033 | Intermediate Similarity | NPD9493 | Approved |
| 0.7969 | Intermediate Similarity | NPD1164 | Approved |
| 0.792 | Intermediate Similarity | NPD2932 | Approved |
| 0.7868 | Intermediate Similarity | NPD2346 | Discontinued |
| 0.7852 | Intermediate Similarity | NPD651 | Clinical (unspecified phase) |
| 0.782 | Intermediate Similarity | NPD943 | Approved |
| 0.7794 | Intermediate Similarity | NPD5406 | Approved |
| 0.7794 | Intermediate Similarity | NPD5405 | Approved |
| 0.7794 | Intermediate Similarity | NPD5408 | Approved |
| 0.7794 | Intermediate Similarity | NPD5404 | Approved |
| 0.7778 | Intermediate Similarity | NPD3019 | Approved |
| 0.7714 | Intermediate Similarity | NPD3300 | Phase 2 |
| 0.7698 | Intermediate Similarity | NPD4196 | Clinical (unspecified phase) |
| 0.7692 | Intermediate Similarity | NPD9261 | Approved |
| 0.7639 | Intermediate Similarity | NPD3226 | Approved |
| 0.7634 | Intermediate Similarity | NPD2798 | Approved |
| 0.7609 | Intermediate Similarity | NPD2344 | Approved |
| 0.7581 | Intermediate Similarity | NPD9281 | Approved |
| 0.7565 | Intermediate Similarity | NPD288 | Approved |
| 0.7536 | Intermediate Similarity | NPD2935 | Discontinued |
| 0.7535 | Intermediate Similarity | NPD7390 | Discontinued |
| 0.7478 | Intermediate Similarity | NPD844 | Approved |
| 0.7464 | Intermediate Similarity | NPD3299 | Clinical (unspecified phase) |
| 0.7464 | Intermediate Similarity | NPD2799 | Discontinued |
| 0.7462 | Intermediate Similarity | NPD9717 | Approved |
| 0.7445 | Intermediate Similarity | NPD1607 | Approved |
| 0.7438 | Intermediate Similarity | NPD9265 | Clinical (unspecified phase) |
| 0.7429 | Intermediate Similarity | NPD970 | Clinical (unspecified phase) |
| 0.7426 | Intermediate Similarity | NPD1240 | Approved |
| 0.7424 | Intermediate Similarity | NPD1203 | Approved |
| 0.7407 | Intermediate Similarity | NPD411 | Approved |
| 0.7407 | Intermediate Similarity | NPD2313 | Discontinued |
| 0.7394 | Intermediate Similarity | NPD2309 | Approved |
| 0.7373 | Intermediate Similarity | NPD289 | Clinical (unspecified phase) |
| 0.7372 | Intermediate Similarity | NPD447 | Suspended |
| 0.7361 | Intermediate Similarity | NPD7422 | Clinical (unspecified phase) |
| 0.7357 | Intermediate Similarity | NPD1471 | Phase 3 |
| 0.7339 | Intermediate Similarity | NPD1609 | Clinical (unspecified phase) |
| 0.7324 | Intermediate Similarity | NPD1878 | Clinical (unspecified phase) |
| 0.7302 | Intermediate Similarity | NPD5951 | Approved |
| 0.7295 | Intermediate Similarity | NPD2342 | Discontinued |
| 0.728 | Intermediate Similarity | NPD7635 | Approved |
| 0.7279 | Intermediate Similarity | NPD3764 | Approved |
| 0.7252 | Intermediate Similarity | NPD1547 | Clinical (unspecified phase) |
| 0.7236 | Intermediate Similarity | NPD6831 | Clinical (unspecified phase) |
| 0.7232 | Intermediate Similarity | NPD650 | Approved |
| 0.7218 | Intermediate Similarity | NPD1283 | Approved |
| 0.7218 | Intermediate Similarity | NPD1876 | Approved |
| 0.7214 | Intermediate Similarity | NPD1510 | Phase 2 |
| 0.7209 | Intermediate Similarity | NPD9545 | Approved |
| 0.719 | Intermediate Similarity | NPD1237 | Approved |
| 0.7177 | Intermediate Similarity | NPD4750 | Phase 3 |
| 0.7176 | Intermediate Similarity | NPD3026 | Approved |
| 0.7176 | Intermediate Similarity | NPD3023 | Approved |
| 0.7155 | Intermediate Similarity | NPD845 | Approved |
| 0.7154 | Intermediate Similarity | NPD1651 | Approved |
| 0.7154 | Intermediate Similarity | NPD3024 | Approved |
| 0.7154 | Intermediate Similarity | NPD3025 | Approved |
| 0.7143 | Intermediate Similarity | NPD1755 | Approved |
| 0.7107 | Intermediate Similarity | NPD1931 | Clinical (unspecified phase) |
| 0.7107 | Intermediate Similarity | NPD1929 | Approved |
| 0.7107 | Intermediate Similarity | NPD1930 | Approved |
| 0.7101 | Intermediate Similarity | NPD520 | Approved |
| 0.7099 | Intermediate Similarity | NPD4626 | Approved |
| 0.7099 | Intermediate Similarity | NPD7163 | Clinical (unspecified phase) |
| 0.7087 | Intermediate Similarity | NPD1241 | Discontinued |
| 0.7083 | Intermediate Similarity | NPD3750 | Approved |
| 0.7083 | Intermediate Similarity | NPD7003 | Approved |
| 0.7068 | Intermediate Similarity | NPD4878 | Approved |
| 0.7059 | Intermediate Similarity | NPD9494 | Approved |
| 0.7049 | Intermediate Similarity | NPD164 | Approved |
| 0.7043 | Intermediate Similarity | NPD9256 | Approved |
| 0.7043 | Intermediate Similarity | NPD9258 | Approved |
| 0.7042 | Intermediate Similarity | NPD1551 | Phase 2 |
| 0.7025 | Intermediate Similarity | NPD1932 | Approved |
| 0.7007 | Intermediate Similarity | NPD2532 | Approved |
| 0.7007 | Intermediate Similarity | NPD2533 | Approved |
| 0.7007 | Intermediate Similarity | NPD2534 | Approved |
| 0.7 | Intermediate Similarity | NPD3020 | Approved |
| 0.6993 | Remote Similarity | NPD2343 | Clinical (unspecified phase) |
| 0.6992 | Remote Similarity | NPD1281 | Approved |
| 0.6987 | Remote Similarity | NPD6232 | Discontinued |
| 0.6984 | Remote Similarity | NPD4141 | Clinical (unspecified phase) |
| 0.6983 | Remote Similarity | NPD7609 | Phase 3 |
| 0.6978 | Remote Similarity | NPD6663 | Approved |
| 0.6974 | Remote Similarity | NPD7819 | Suspended |
| 0.6966 | Remote Similarity | NPD8166 | Discontinued |
| 0.6962 | Remote Similarity | NPD7473 | Discontinued |
| 0.6949 | Remote Similarity | NPD2934 | Approved |
| 0.6949 | Remote Similarity | NPD1432 | Clinical (unspecified phase) |
| 0.6949 | Remote Similarity | NPD2933 | Approved |
| 0.6944 | Remote Similarity | NPD1549 | Phase 2 |
| 0.694 | Remote Similarity | NPD9269 | Phase 2 |
| 0.694 | Remote Similarity | NPD3972 | Approved |
| 0.6939 | Remote Similarity | NPD1511 | Approved |
| 0.6934 | Remote Similarity | NPD5736 | Approved |
| 0.6923 | Remote Similarity | NPD7631 | Approved |
| 0.6897 | Remote Similarity | NPD2800 | Approved |
| 0.6894 | Remote Similarity | NPD9268 | Approved |
| 0.6892 | Remote Similarity | NPD4378 | Clinical (unspecified phase) |
| 0.6891 | Remote Similarity | NPD2859 | Approved |
| 0.6891 | Remote Similarity | NPD1809 | Phase 2 |
| 0.6891 | Remote Similarity | NPD2860 | Approved |
| 0.6884 | Remote Similarity | NPD6832 | Phase 2 |
| 0.6879 | Remote Similarity | NPD1899 | Clinical (unspecified phase) |
| 0.6875 | Remote Similarity | NPD1552 | Clinical (unspecified phase) |
| 0.6875 | Remote Similarity | NPD1550 | Clinical (unspecified phase) |
| 0.686 | Remote Similarity | NPD2066 | Phase 3 |
| 0.6853 | Remote Similarity | NPD4308 | Phase 3 |
| 0.6849 | Remote Similarity | NPD6398 | Clinical (unspecified phase) |
| 0.6846 | Remote Similarity | NPD1512 | Approved |
| 0.6842 | Remote Similarity | NPD3412 | Clinical (unspecified phase) |
| 0.6825 | Remote Similarity | NPD2329 | Discontinued |
| 0.6825 | Remote Similarity | NPD2182 | Approved |
| 0.6821 | Remote Similarity | NPD5808 | Clinical (unspecified phase) |
| 0.6818 | Remote Similarity | NPD2296 | Approved |
| 0.6818 | Remote Similarity | NPD3091 | Approved |
| 0.6815 | Remote Similarity | NPD1608 | Approved |
| 0.6788 | Remote Similarity | NPD2797 | Approved |
| 0.6779 | Remote Similarity | NPD7410 | Clinical (unspecified phase) |
| 0.6774 | Remote Similarity | NPD3882 | Suspended |
| 0.6772 | Remote Similarity | NPD3021 | Approved |
| 0.6772 | Remote Similarity | NPD3022 | Approved |
| 0.6772 | Remote Similarity | NPD1792 | Phase 2 |
| 0.6769 | Remote Similarity | NPD497 | Approved |
| 0.6767 | Remote Similarity | NPD5691 | Approved |
| 0.6761 | Remote Similarity | NPD230 | Phase 1 |
| 0.6754 | Remote Similarity | NPD111 | Approved |
| 0.6753 | Remote Similarity | NPD2393 | Clinical (unspecified phase) |
| 0.6753 | Remote Similarity | NPD2801 | Approved |
| 0.6748 | Remote Similarity | NPD846 | Approved |
| 0.6748 | Remote Similarity | NPD5031 | Approved |
| 0.6748 | Remote Similarity | NPD940 | Approved |
| 0.6748 | Remote Similarity | NPD5027 | Approved |
| 0.6748 | Remote Similarity | NPD5029 | Approved |
| 0.6739 | Remote Similarity | NPD1019 | Discontinued |
| 0.6736 | Remote Similarity | NPD3748 | Approved |
| 0.6733 | Remote Similarity | NPD6273 | Approved |
| 0.6731 | Remote Similarity | NPD3749 | Approved |
| 0.6723 | Remote Similarity | NPD1202 | Approved |
| 0.6719 | Remote Similarity | NPD2181 | Clinical (unspecified phase) |
| 0.6718 | Remote Similarity | NPD256 | Approved |
| 0.6718 | Remote Similarity | NPD255 | Approved |
| 0.6716 | Remote Similarity | NPD4059 | Approved |
| 0.6711 | Remote Similarity | NPD2651 | Approved |
| 0.6711 | Remote Similarity | NPD2649 | Approved |
| 0.6709 | Remote Similarity | NPD6959 | Discontinued |
| 0.6707 | Remote Similarity | NPD5034 | Approved |
| 0.6707 | Remote Similarity | NPD36 | Approved |
| 0.6707 | Remote Similarity | NPD4954 | Approved |
| 0.6707 | Remote Similarity | NPD5028 | Approved |
| 0.6707 | Remote Similarity | NPD4955 | Approved |
| 0.6707 | Remote Similarity | NPD5026 | Approved |
| 0.6696 | Remote Similarity | NPD9257 | Approved |
| 0.6696 | Remote Similarity | NPD9259 | Approved |
| 0.6692 | Remote Similarity | NPD2226 | Clinical (unspecified phase) |
| 0.6692 | Remote Similarity | NPD498 | Approved |
| 0.6692 | Remote Similarity | NPD495 | Approved |
| 0.6692 | Remote Similarity | NPD1759 | Phase 1 |
| 0.6692 | Remote Similarity | NPD496 | Approved |
| 0.669 | Remote Similarity | NPD6100 | Approved |
| 0.669 | Remote Similarity | NPD6099 | Approved |
| 0.669 | Remote Similarity | NPD2796 | Approved |
| 0.6688 | Remote Similarity | NPD1934 | Approved |
| 0.6688 | Remote Similarity | NPD6844 | Discontinued |
| 0.6667 | Remote Similarity | NPD1242 | Phase 1 |
| 0.6667 | Remote Similarity | NPD688 | Clinical (unspecified phase) |
| 0.6667 | Remote Similarity | NPD4879 | Approved |
| 0.6667 | Remote Similarity | NPD3268 | Approved |
| 0.6667 | Remote Similarity | NPD4380 | Phase 2 |
| 0.6667 | Remote Similarity | NPD7421 | Clinical (unspecified phase) |
| 0.6667 | Remote Similarity | NPD1445 | Approved |
| 0.6667 | Remote Similarity | NPD1444 | Approved |
| 0.6647 | Remote Similarity | NPD8150 | Discontinued |
| 0.6646 | Remote Similarity | NPD5030 | Phase 2 |
| 0.6644 | Remote Similarity | NPD6884 | Clinical (unspecified phase) |
| 0.6642 | Remote Similarity | NPD3600 | Clinical (unspecified phase) |
| 0.6642 | Remote Similarity | NPD4379 | Clinical (unspecified phase) |
| 0.6642 | Remote Similarity | NPD182 | Clinical (unspecified phase) |
| 0.6622 | Remote Similarity | NPD1196 | Approved |
| 0.6618 | Remote Similarity | NPD3092 | Approved |
| 0.6617 | Remote Similarity | NPD1758 | Phase 1 |
| 0.6603 | Remote Similarity | NPD8443 | Clinical (unspecified phase) |
| 0.6601 | Remote Similarity | NPD7458 | Discontinued |
| 0.66 | Remote Similarity | NPD1543 | Discontinued |
| 0.6597 | Remote Similarity | NPD6651 | Approved |
| 0.6596 | Remote Similarity | NPD7008 | Discontinued |
| 0.6596 | Remote Similarity | NPD5952 | Clinical (unspecified phase) |
| 0.6593 | Remote Similarity | NPD2286 | Discontinued |
| 0.6593 | Remote Similarity | NPD17 | Approved |
| 0.6593 | Remote Similarity | NPD1778 | Approved |
 Bioactivity similarity: Similar Natural Products in NPASS
Bioactivity similarity: Similar Natural Products in NPASSBioactivity similarity was calculated based on bioactivity descriptors of compounds. The bioactivity descriptors were calculated by a recently developed AI algorithm Chemical Checker (CC) [Nature Biotechnology, 38:1087–1096, 2020; Nature Communications, 12:3932, 2021], which evaluated bioactivity similarities at five levels:
☉ A: chemistry similarity;
☉ B: biological targets similarity;
☉ C: networks similarity;
☉ D: cell-based bioactivity similarity;
☉ E: similarity based on clinical data.
Those 5 categories of CC bioactivity descriptors were calculated and then subjected to manifold projection using UMAP algorithm, to project all NPs on a 2-Dimensional space. The current NP was highlighted with a small circle in the 2-D map. Below figures: left-to-right, A-to-E.