| Molecular Weight: | 302.04 |
| Volume: | 282.767 |
| LogP: | 2.079 |
| LogD: | 1.942 |
| LogS: | -3.497 |
| # Rotatable Bonds: | 1 |
| TPSA: | 131.36 |
| # H-Bond Aceptor: | 7 |
| # H-Bond Donor: | 5 |
| # Rings: | 3 |
| # Heavy Atoms: | 7 |
| QED Drug-Likeness Score: | 0.464 |
| Synthetic Accessibility Score: | 2.613 |
| Fsp3: | 0.0 |
| Lipinski Rule-of-5: | Accepted |
| Pfizer Rule: | Accepted |
| GSK Rule: | Accepted |
| BMS Rule: | 1 |
| Golden Triangle Rule: | Accepted |
| Chelating Alert: | 1 |
| PAINS Alert: | 0 |
| Caco-2 Permeability: | -5.182 |
| MDCK Permeability: | 7.003451173659414e-06 |
| Pgp-inhibitor: | 0.005 |
| Pgp-substrate: | 0.007 |
| Human Intestinal Absorption (HIA): | 0.012 |
| 20% Bioavailability (F20%): | 0.939 |
| 30% Bioavailability (F30%): | 0.998 |
| Blood-Brain-Barrier Penetration (BBB): | 0.011 |
| Plasma Protein Binding (PPB): | 96.2477035522461% |
| Volume Distribution (VD): | 0.551 |
| Pgp-substrate: | 6.231308937072754% |
| CYP1A2-inhibitor: | 0.953 |
| CYP1A2-substrate: | 0.116 |
| CYP2C19-inhibitor: | 0.087 |
| CYP2C19-substrate: | 0.041 |
| CYP2C9-inhibitor: | 0.636 |
| CYP2C9-substrate: | 0.826 |
| CYP2D6-inhibitor: | 0.587 |
| CYP2D6-substrate: | 0.242 |
| CYP3A4-inhibitor: | 0.618 |
| CYP3A4-substrate: | 0.051 |
| Clearance (CL): | 7.912 |
| Half-life (T1/2): | 0.931 |
| hERG Blockers: | 0.157 |
| Human Hepatotoxicity (H-HT): | 0.067 |
| Drug-inuced Liver Injury (DILI): | 0.979 |
| AMES Toxicity: | 0.616 |
| Rat Oral Acute Toxicity: | 0.053 |
| Maximum Recommended Daily Dose: | 0.386 |
| Skin Sensitization: | 0.87 |
| Carcinogencity: | 0.035 |
| Eye Corrosion: | 0.009 |
| Eye Irritation: | 0.931 |
| Respiratory Toxicity: | 0.057 |
 Natural Product: NPC216361
Natural Product: NPC216361| Natural Product ID: | NPC216361 |
| Common Name*: | Morin |
| IUPAC Name: | 2-(2,4-dihydroxyphenyl)-3,5,7-trihydroxychromen-4-one |
| Synonyms: | NSC-19801 |
| Standard InCHIKey: | YXOLAZRVSSWPPT-UHFFFAOYSA-N |
| Standard InCHI: | InChI=1S/C15H10O7/c16-6-1-2-8(9(18)3-6)15-14(21)13(20)12-10(19)4-7(17)5-11(12)22-15/h1-5,16-19,21H |
| SMILES: | Oc1ccc(c(c1)O)c1oc2cc(O)cc(c2c(=O)c1O)O |
| Synthetic Gene Cluster: | n.a. |
| ChEMBL Identifier: | CHEMBL28626 |
| PubChem CID: |
5281670 |
| Chemical Classification**: |
|
*Note: the InCHIKey will be temporarily assigned as the "Common Name" if no IUPAC name or alternative short name is available.
**Note: the Chemical Classification was calculated by NPClassifier Version 1.5. Reference: PMID:34662515.
 Species Source
Species Source| Organism ID | Organism Name | Taxonomy Level | Family | SuperKingdom | Isolation Part | Collection Location | Collection Time | Reference |
|---|---|---|---|---|---|---|---|---|
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | DOI[10.1016/j.phytochem.2005.10.025] |
| NPO3806 | Garcinia morella | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | DOI[10.1016/S0040-4039(00)90246-6] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Fruit | n.a. | n.a. | DOI[10.1016/S0963-9969(99)00095-2] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Leaves (mother plant) | Suwon, Korea | 2011-Sep | DOI[10.7235/HORT.2013.12200] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Roots (runner plant) | Suwon, Korea | 2011-Sep | DOI[10.7235/HORT.2013.12200] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Fruits | Suwon, Korea | 2012-Nov | DOI[10.7235/HORT.2013.12200] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Leaves | Suwon, Korea | DOI[10.7235/HORT.2013.12200] | |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Runners | Suwon, Korea | 2011-Sep | DOI[10.7235/HORT.2013.12200] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Leaves (runner plant) | Suwon, Korea | 2011-Sep | DOI[10.7235/HORT.2013.12200] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Roots (mother plant) | Suwon, Korea | 2011-Sep | DOI[10.7235/HORT.2013.12200] |
| NPO7230 | Artocarpus champeden | Species | Moraceae | Eukaryota | Roots | n.a. | n.a. |
PMID[10217722] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | fruit body | n.a. |
PMID[11520245] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[11908994] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[12045343] |
| NPO16949 | Aster oharai | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[12193011] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | stems | n.a. | n.a. |
PMID[12932126] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[1453178] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[14695801] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Fruits | n.a. | n.a. |
PMID[15568772] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[15787460] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[15787460] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[16203143] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[16919944] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | root bark | n.a. | n.a. |
PMID[17608532] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[1791484] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[17950599] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | rhizomes | n.a. | n.a. |
PMID[18177011] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | root | n.a. |
PMID[1823989] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | spore | n.a. |
PMID[18827358] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | fruits | n.a. | n.a. |
PMID[19113968] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | Roots | n.a. | n.a. |
PMID[19191562] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[19271742] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[19280148] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[19285413] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[19660948] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | fruiting bodies | Nagano, Japan | 2008-MAY |
PMID[20039640] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[20384295] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[20590154] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | twigs | n.a. | n.a. |
PMID[20681570] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | fruiting bodies | n.a. | n.a. |
PMID[20702092] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[20815207] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[21319773] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[21319773] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | bark | n.a. |
PMID[21319773] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | leaf | n.a. |
PMID[21319773] |
| NPO28834 | Anabaena minutissima | Species | 0stocaceae | Bacteria | n.a. | n.a. | n.a. |
PMID[21699148] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | fruiting bodies | n.a. | n.a. |
PMID[21924611] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[22014750] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | fruit body | n.a. |
PMID[22044278] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | fruiting body | n.a. | n.a. |
PMID[22047696] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | leaf | n.a. |
PMID[22207282] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[23806866] |
| NPO22730 | Indigofera spicata | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[23895019] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | root bark | University of Veterinary and Pharmaceutical Sciences Brno (UVPS Brno), Brno, Czech Republic | 2011-APR |
PMID[24901948] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | Konya, Turkey | 2007-APR |
PMID[24901948] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | Roots | n.a. | n.a. |
PMID[25322455] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | leaves | Chungbuk, Korea | n.a. |
PMID[25935644] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | root | n.a. |
PMID[25981820] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[26222905] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | root barks | purchased from the local herbal market, Chungbuk, Korea | 2012-JUL |
PMID[26227773] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | latex | n.a. |
PMID[27294120] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | Root barks | n.a. | n.a. |
PMID[3097265] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[31747281] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[8064299] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | rhizome | n.a. |
PMID[8064299] |
| NPO30745 | Chrysanthemum morifolium | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[8158164] |
| NPO30745 | Chrysanthemum morifolium | Species | Asteraceae | Eukaryota | n.a. | inflorescence | n.a. |
PMID[8158164] |
| NPO40537 | Koelreuteria henryi | Species | Sapindaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[8350096] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. |
PMID[8594148] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | Twigs and leaves | Taichung, Taiwan | 1993-Aug |
PMID[9322359] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | fruit body | n.a. |
PMID[9635497] |
| NPO28241 | Agelas conifera | Species | Agelasidae | Eukaryota | n.a. | n.a. | n.a. |
PMID[9784176] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | Plant | n.a. | n.a. | Database[FooDB] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | Resin, Exudate, Sap | n.a. | n.a. | Database[FooDB] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | Rhizome | n.a. | n.a. | Database[FooDB] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | Root | n.a. | n.a. | Database[FooDB] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Fruits | n.a. | Database[FooDB] | |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | n.a. | n.a. | Database[FooDB] | |
| NPO17655 | Heliotropium amplexicaule | Species | Heliotropiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO31031 | Artocarpus integrifolia | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO16511 | Iberis amara | Species | Brassicaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO14125 | Corydalis ochotensis | Species | Papaveraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO31003 | Moschus sifanicus | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO10437 | Periploca calophylla | Species | Apocynaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO28498 | Fusarium roseum | Species | Nectriaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO6326 | Morus serrata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO30745 | Chrysanthemum morifolium | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO67 | Fructus mori | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO3806 | Garcinia morella | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO22978 | Morus tinctoria | Species | Sulidae | Eukaryota | n.a. | n.a. | n.a. | Database[HerDing] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | Fruits | n.a. | Database[Phenol-Explorer] | |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | Database[Phenol-Explorer] | |
| NPO10437 | Periploca calophylla | Species | Apocynaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO4183 | Radix cudraniae | Species | Lymnaeidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO463 | Moschus berezovskii | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO2646 | Moschus moschiferus | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO22978 | Morus tinctoria | Species | Sulidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO8300 | Apis mellifera ligustica | Species | Apidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO16511 | Iberis amara | Species | Brassicaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO8964 | Herba lagotidis | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO67 | Fructus mori | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO19905 | Cortex mori radicis praeparata | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO7230 | Artocarpus champeden | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO12246 | Moschus chrysogaster | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO28498 | Fusarium roseum | Species | Nectriaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO17655 | Heliotropium amplexicaule | Species | Heliotropiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO14125 | Corydalis ochotensis | Species | Papaveraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO6326 | Morus serrata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO3806 | Garcinia morella | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCMID] |
| NPO30745 | Chrysanthemum morifolium | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO2646 | Moschus moschiferus | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO30804 | Cudrania tricuspidata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO463 | Moschus berezovskii | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO3806 | Garcinia morella | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO31003 | Moschus sifanicus | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO16511 | Iberis amara | Species | Brassicaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TCM_Taiwan] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO16609 | Ganoderma sinense | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO20553 | Zingiber officinale | Species | Zingiberaceae | Eukaryota | n.a. | n.a. | n.a. | Database[TM-MC] |
| NPO26351 | Agastache rugosa | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO25269 | Ganoderma lucidum | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16735 | Sophora tometosa | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17245 | Prumnopitys ferruginoides | Species | Podocarpaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16511 | Iberis amara | Species | Brassicaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28834 | Anabaena minutissima | Species | 0stocaceae | Bacteria | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28498 | Fusarium roseum | Species | Nectriaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO2646 | Moschus moschiferus | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO9860.2 | Pinus brutia var. pityusa | Varieties | Pinaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22978 | Morus tinctoria | Species | Sulidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16949 | Aster oharai | Species | Asteraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO10437 | Periploca calophylla | Species | Apocynaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO6958 | Pseudaegle trifoliata | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28546 | Entomophthora virulenta | Species | Entomophthoraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO463 | Moschus berezovskii | Species | Moschidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO14880 | Ephydatia syriaca | Species | Spongillidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28199 | Sargassum thunbergii | Species | Sargassaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28815 | Phycopsis terpnis | Species | Bubaridae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO16609 | Ganoderma sinense | Species | Ganodermataceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO3374 | Pestalotiopsis guepinii | Species | Sporocadaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO3806 | Garcinia morella | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO29081 | Plantago bellardi | Species | Plantaginaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO24162 | Erysimum canescens | Species | Brassicaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO6326 | Morus serrata | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO22730 | Indigofera spicata | Species | Fabaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO11568 | Dendroctonus ponderosae | Species | Curculionidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO10339 | Fragaria x ananassa | Species | Rosaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO11569 | Hypsizygus ulmarius | Species | Lyophyllaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO5049 | Garcinia multiflora | Species | Clusiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28629 | Hedeoma multiflora | Species | Lamiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28265 | Morus alba | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO14125 | Corydalis ochotensis | Species | Papaveraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO18109 | Thorecta marginalis | Species | Thorectidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO28241 | Agelas conifera | Species | Agelasidae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO17655 | Heliotropium amplexicaule | Species | Heliotropiaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO7230 | Artocarpus champeden | Species | Moraceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
| NPO8527 | Holboellia fargesi | Species | Lardizabalaceae | Eukaryota | n.a. | n.a. | n.a. | Database[UNPD] |
☑ Note for Reference:
In addition to directly collecting NP source organism data from primary literature (where reference will provided as NCBI PMID or DOI links), NPASS also integrated them from below databases:
☉ UNPD: Universal Natural Products Database [PMID: 23638153].
☉ StreptomeDB: a database of streptomycetes natural products [PMID: 33051671].
☉ TM-MC: a database of medicinal materials and chemical compounds in Northeast Asian traditional medicine [PMID: 26156871].
☉ TCM@Taiwan: a Traditional Chinese Medicine database [PMID: 21253603].
☉ TCMID: a Traditional Chinese Medicine database [PMID: 29106634].
☉ TCMSP: The traditional Chinese medicine systems pharmacology database and analysis platform [PMID: 24735618].
☉ HerDing: a herb recommendation system to treat diseases using genes and chemicals [PMID: 26980517].
☉ MetaboLights: a metabolomics database [PMID: 27010336].
☉ FooDB: a database of constituents, chemistry and biology of food species [www.foodb.ca].
 NP Quantity Composition/Concentration
NP Quantity Composition/Concentration| Organism ID | NP ID | Organism Material Preparation | Organism Part | NP Quantity (Standard) | NP Quantity (Minimum) | NP Quantity (Maximum) | Quantity Unit | Reference |
|---|---|---|---|---|---|---|---|---|
| NPO10339 | NPC216361 | Raw | n.a. | 0.060000002 | n.a. | n.a. | mg/100g | Database [PHENOL EXPLORER] |
| NPO10339 | NPC216361 | n.a. | Fruits | 0.06 | n.a. | n.a. | mg/100g of FW | Database [Phenol-Explorer] |
☑ Note for Reference:
In addition to directly collecting NP quantitative data from primary literature (where reference will provided as NCBI PMID or DOI links), NPASS also integrated NP quantitative records for specific NP domains (e.g., NPS from foods or herbs) from domain-specific databases. These databases include:
☉ DUKE: Dr. Duke's Phytochemical and Ethnobotanical Databases.
☉ PHENOL EXPLORER: is the first comprehensive database on polyphenol content in foods [PMID: 24103452], its homepage can be accessed at here.
☉ FooDB: a database of constituents, chemistry and biology of food species [www.foodb.ca].
 Biological Activity
Biological Activity| Target ID | Target Type | Target Name | Target Organism | Activity Type | Activity Relation | Value | Unit | Reference |
|---|---|---|---|---|---|---|---|---|
| NPT1315 | Individual Protein | Adenosine A1 receptor | Rattus norvegicus | Ki | = | 13800.0 | nM | PMID[485857] |
| NPT218 | Individual Protein | Adenosine A3 receptor | Homo sapiens | Ki | = | 34000.0 | nM | PMID[485857] |
| NPT12 | Individual Protein | Human immunodeficiency virus type 1 integrase | Human immunodeficiency virus 1 | IC50 | = | 76500.0 | nM | PMID[485858] |
| NPT12 | Individual Protein | Human immunodeficiency virus type 1 integrase | Human immunodeficiency virus 1 | IC50 | = | 31700.0 | nM | PMID[485858] |
| NPT1566 | Individual Protein | Glyoxalase I | Homo sapiens | Ki | = | 30199.52 | nM | PMID[485859] |
| NPT1568 | Individual Protein | Trypsin I | Bos taurus | IC50 | = | 110800.0 | nM | PMID[485860] |
| NPT1332 | Individual Protein | Fatty acid synthase | Plasmodium falciparum | IC50 | = | 8000.0 | nM | PMID[485861] |
| NPT1333 | Individual Protein | Enoyl-acyl-carrier protein reductase | Plasmodium falciparum | IC50 | = | 5000.0 | nM | PMID[485861] |
| NPT1334 | Individual Protein | 3-oxoacyl-acyl-carrier protein reductase | Plasmodium falciparum | IC50 | = | 2300.0 | nM | PMID[485861] |
| NPT1265 | Individual Protein | Serine/threonine-protein kinase PIM1 | Homo sapiens | -Log IC50 | = | -0.43 | uM | PMID[485863] |
| NPT5233 | Individual Protein | Solute carrier family 22 member 12 | Rattus norvegicus | IC50 | = | 18000.0 | nM | PMID[485864] |
| NPT5234 | Individual Protein | Solute carrier family 22 member 12 | Homo sapiens | IC50 | = | 2000.0 | nM | PMID[485864] |
| NPT5234 | Individual Protein | Solute carrier family 22 member 12 | Homo sapiens | Ki | = | 5740.0 | nM | PMID[485864] |
| NPT1821 | Individual Protein | Sortase | Staphylococcus aureus | IC50 | = | 37390.0 | nM | PMID[485867] |
| NPT1460 | Cell Line | L929 | Mus musculus | EC50 | = | 120000.0 | nM | PMID[485870] |
| NPT1460 | Cell Line | L929 | Mus musculus | EC50 | = | 115000.0 | nM | PMID[485870] |
| NPT612 | Individual Protein | Canalicular multispecific organic anion transporter 1 | Homo sapiens | Activity | = | 50.0 | % | PMID[485874] |
| NPT1044 | Individual Protein | DNA topoisomerase II alpha | Homo sapiens | Activity | < | 50.0 | % | PMID[485875] |
| NPT759 | Cell Line | H9 | Homo sapiens | IC50 | > | 331000.0 | nM | PMID[485877] |
| NPT1044 | Individual Protein | DNA topoisomerase II alpha | Homo sapiens | IC50 | = | 40.8 | ug.mL-1 | PMID[485875] |
| NPT163 | Individual Protein | Nuclear factor NF-kappa-B p105 subunit | Homo sapiens | Potency | = | 10000.0 | nM | PMID[485888] |
| NPT47 | Individual Protein | ATP-dependent DNA helicase Q1 | Homo sapiens | Potency | = | 7079.5 | nM | PMID[485887] |
| NPT47 | Individual Protein | ATP-dependent DNA helicase Q1 | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485885] |
| NPT2741 | Individual Protein | Pyruvate kinase | Leishmania mexicana | Potency | = | 19952.6 | nM | PMID[485888] |
| NPT47 | Individual Protein | ATP-dependent DNA helicase Q1 | Homo sapiens | Potency | = | 28183.8 | nM | PMID[485888] |
| NPT47 | Individual Protein | ATP-dependent DNA helicase Q1 | Homo sapiens | Potency | = | 39810.7 | nM | PMID[485886] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 35481.3 | nM | PMID[485885] |
| NPT149 | Individual Protein | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Homo sapiens | Potency | = | 12589.3 | nM | PMID[485888] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 25118.9 | nM | PMID[485887] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 28183.8 | nM | PMID[485888] |
| NPT48 | Individual Protein | Lysine-specific demethylase 4D-like | Homo sapiens | Potency | = | 39810.7 | nM | PMID[485886] |
| NPT1038 | Individual Protein | Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 35481.3 | nM | PMID[485889] |
| NPT1226 | Individual Protein | Caspase-7 | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485888] |
| NPT50 | Individual Protein | Tyrosyl-DNA phosphodiesterase 1 | Homo sapiens | Potency | = | 199.5 | nM | PMID[485887] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 11220.2 | nM | PMID[485885] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 28183.8 | nM | PMID[485887] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485886] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 14125.4 | nM | PMID[485888] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485887] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 12589.3 | nM | PMID[485885] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 12589.3 | nM | PMID[485888] |
| NPT248 | Individual Protein | Estrogen receptor beta | Homo sapiens | IC50 | = | 2927.18 | nM | PMID[485888] |
| NPT51 | Individual Protein | Microtubule-associated protein tau | Homo sapiens | Potency | = | 112.2 | nM | PMID[485886] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | = | 5011.9 | nM | PMID[485885] |
| NPT57 | Individual Protein | Induced myeloid leukemia cell differentiation protein Mcl-1 | Homo sapiens | IC50 | = | 2988.26 | nM | PMID[485888] |
| NPT58 | Individual Protein | Bloom syndrome protein | Homo sapiens | Potency | = | 19952.6 | nM | PMID[485885] |
| NPT151 | Individual Protein | 15-hydroxyprostaglandin dehydrogenase [NAD+] | Homo sapiens | Potency | = | 31622.8 | nM | PMID[485885] |
| NPT151 | Individual Protein | 15-hydroxyprostaglandin dehydrogenase [NAD+] | Homo sapiens | Potency | = | 35481.3 | nM | PMID[485887] |
| NPT277 | Individual Protein | Caspase-1 | Homo sapiens | Potency | = | 39810.7 | nM | PMID[485888] |
| NPT149 | Individual Protein | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Homo sapiens | Potency | = | 12589.3 | nM | PMID[485891] |
| NPT539 | Individual Protein | Cellular tumor antigen p53 | Homo sapiens | Potency | = | 3981.1 | nM | PMID[485888] |
| NPT149 | Individual Protein | Endoplasmic reticulum-associated amyloid beta-peptide-binding protein | Homo sapiens | Potency | = | 10000.0 | nM | PMID[485888] |
| NPT60 | Individual Protein | Lysosomal alpha-glucosidase | Homo sapiens | Potency | = | 3162.3 | nM | PMID[485888] |
| NPT59 | Individual Protein | DNA polymerase beta | Homo sapiens | Potency | = | 562.3 | nM | PMID[485885] |
| NPT109 | Individual Protein | Cytochrome P450 3A4 | Homo sapiens | Potency | = | 31622.8 | nM | PMID[485888] |
| NPT59 | Individual Protein | DNA polymerase beta | Homo sapiens | Potency | = | 4466.8 | nM | PMID[485888] |
| NPT45 | Individual Protein | Ubiquitin carboxyl-terminal hydrolase 2 | Homo sapiens | Potency | = | 18492.7 | nM | PMID[485890] |
| NPT212 | Individual Protein | Cytochrome P450 2C9 | Homo sapiens | Potency | = | 10000.0 | nM | PMID[485891] |
| NPT1422 | Individual Protein | ATP-binding cassette sub-family G member 2 | Homo sapiens | IC50 | = | 21000.0 | nM | PMID[485892] |
| NPT1422 | Individual Protein | ATP-binding cassette sub-family G member 2 | Homo sapiens | IC50 | = | 49000.0 | nM | PMID[485892] |
| NPT443 | Individual Protein | Histone acetyltransferase GCN5 | Homo sapiens | Potency | n.a. | 15848.9 | nM | PMID[485890] |
| NPT62 | Individual Protein | 6-phospho-1-fructokinase | Trypanosoma brucei | Potency | n.a. | 23934.1 | nM | PMID[485888] |
| NPT242 | Individual Protein | Dopamine D1 receptor | Homo sapiens | Potency | n.a. | 103.2 | nM | PMID[485890] |
| NPT1623 | Individual Protein | DNA polymerase III | Bacillus subtilis (strain 168) | Potency | n.a. | 18887.6 | nM | PMID[485890] |
| NPT5 | Individual Protein | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 | Homo sapiens | Potency | n.a. | 39810.7 | nM | PMID[485890] |
| NPT5 | Individual Protein | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 | Homo sapiens | Potency | n.a. | 39810.7 | nM | PMID[485888] |
| NPT62 | Individual Protein | 6-phospho-1-fructokinase | Trypanosoma brucei | Potency | n.a. | 6012.0 | nM | PMID[485890] |
| NPT135 | Individual Protein | Chromobox protein homolog 1 | Homo sapiens | Potency | n.a. | 39810.7 | nM | PMID[485890] |
| NPT534 | Individual Protein | Phosphoglycerate kinase, glycosomal | Trypanosoma brucei brucei | Potency | n.a. | 23934.1 | nM | PMID[485890] |
| NPT63 | Individual Protein | Bromodomain adjacent to zinc finger domain protein 2B | Homo sapiens | Potency | n.a. | 50118.7 | nM | PMID[485888] |
| NPT199 | Individual Protein | DNA polymerase kappa | Homo sapiens | Potency | n.a. | 3162.3 | nM | PMID[485895] |
| NPT199 | Individual Protein | DNA polymerase kappa | Homo sapiens | Potency | n.a. | 4466.8 | nM | PMID[485896] |
| NPT199 | Individual Protein | DNA polymerase kappa | Homo sapiens | Potency | n.a. | 22387.2 | nM | PMID[485894] |
| NPT199 | Individual Protein | DNA polymerase kappa | Homo sapiens | Potency | n.a. | 4228.4 | nM | PMID[485890] |
| NPT71 | Cell Line | HEK293 | Homo sapiens | Potency | n.a. | 3.1 | nM | PMID[485890] |
| NPT71 | Cell Line | HEK293 | Homo sapiens | Potency | n.a. | 12589.3 | nM | PMID[485885] |
| NPT444 | Individual Protein | Ubiquitin carboxyl-terminal hydrolase 1 | Homo sapiens | Potency | n.a. | 50118.7 | nM | PMID[485890] |
| NPT367 | Cell Line | MDA-N | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT368 | Cell Line | SN12C | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT369 | Cell Line | ACHN | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT370 | Cell Line | NCI-H23 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT371 | Cell Line | UO-31 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT116 | Cell Line | HL-60 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT372 | Cell Line | HOP-92 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT90 | Cell Line | DU-145 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT374 | Cell Line | SF-539 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT373 | Cell Line | SK-MEL-5 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT375 | Cell Line | Malme-3M | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT111 | Cell Line | K562 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT377 | Cell Line | OVCAR-3 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT379 | Cell Line | HOP-62 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT378 | Cell Line | NCI/ADR-RES | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT381 | Cell Line | OVCAR-8 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT380 | Cell Line | U-251 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT382 | Cell Line | OVCAR-5 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT383 | Cell Line | SNB-19 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT82 | Cell Line | MDA-MB-231 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT384 | Cell Line | TK-10 | Homo sapiens | GI50 | n.a. | 33419.5 | nM | PMID[485897] |
| NPT323 | Cell Line | SW-620 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT455 | Cell Line | NCI-H522 | Homo sapiens | GI50 | n.a. | 10000.0 | nM | PMID[485897] |
| NPT386 | Cell Line | KM12 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT387 | Cell Line | M14 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT388 | Cell Line | NCI-H322M | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT456 | Cell Line | OVCAR-4 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT390 | Cell Line | LOX IMVI | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT147 | Cell Line | SK-MEL-2 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT391 | Cell Line | HCC 2998 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT81 | Cell Line | A549 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT392 | Cell Line | SNB-75 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT393 | Cell Line | HCT-116 | Homo sapiens | GI50 | n.a. | 86099.38 | nM | PMID[485897] |
| NPT148 | Cell Line | HCT-15 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT394 | Cell Line | EKVX | Homo sapiens | GI50 | n.a. | 30269.13 | nM | PMID[485897] |
| NPT395 | Cell Line | SF-268 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT306 | Cell Line | PC-3 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT83 | Cell Line | MCF7 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT396 | Cell Line | T47D | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT397 | Cell Line | NCI-H460 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT308 | Cell Line | CAKI-1 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT400 | Cell Line | MDA-MB-435 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT458 | Cell Line | IGROV-1 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT399 | Cell Line | SF-295 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT401 | Cell Line | 786-0 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT402 | Cell Line | Hs-578T | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT403 | Cell Line | UACC-257 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT404 | Cell Line | CCRF-CEM | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT139 | Cell Line | HT-29 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT405 | Cell Line | NCI-H226 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT170 | Cell Line | SK-MEL-28 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT407 | Cell Line | COLO 205 | Homo sapiens | GI50 | n.a. | 100000.0 | nM | PMID[485897] |
| NPT1626 | Individual Protein | Protein E6 | Human papillomavirus type 16 | IC50 | = | 4000.0 | nM | PMID[485898] |
| NPT1627 | Individual Protein | G-protein coupled receptor 35 | Homo sapiens | EC50 | = | 1330.0 | nM | PMID[485900] |
| NPT1627 | Individual Protein | G-protein coupled receptor 35 | Homo sapiens | EC50 | = | 11200.0 | nM | PMID[485900] |
| NPT1627 | Individual Protein | G-protein coupled receptor 35 | Homo sapiens | IC50 | = | 1600.0 | nM | PMID[485900] |
| NPT1627 | Individual Protein | G-protein coupled receptor 35 | Homo sapiens | Efficacy | = | 284.0 | picometer | PMID[485900] |
| NPT1627 | Individual Protein | G-protein coupled receptor 35 | Homo sapiens | Ratio | = | 9.8 | n.a. | PMID[485900] |
| NPT1594 | Individual Protein | Multidrug resistance-associated protein 1 | Homo sapiens | Activity | = | 546.0 | % | PMID[485902] |
| NPT1594 | Individual Protein | Multidrug resistance-associated protein 1 | Homo sapiens | Activity | = | 553.0 | % | PMID[485902] |
| NPT1633 | Individual Protein | Low molecular weight phosphotyrosine protein phosphatase | Bos taurus | IC50 | = | 10000.0 | nM | PMID[485903] |
| NPT1633 | Individual Protein | Low molecular weight phosphotyrosine protein phosphatase | Bos taurus | Inhibition | = | 35.0 | % | PMID[485903] |
| NPT622 | Individual Protein | Solute carrier organic anion transporter family member 2B1 | Homo sapiens | Inhibition | = | 71.3 | % | PMID[485905] |
| NPT66 | Individual Protein | Acetylcholinesterase | Electrophorus electricus | Inhibition | = | 18.1 | % | PMID[485906] |
| NPT1449 | Individual Protein | Tyrosine-protein kinase SYK | Homo sapiens | EC50 | > | 300000.0 | nM | PMID[485907] |
| NPT1449 | Individual Protein | Tyrosine-protein kinase SYK | Homo sapiens | IC50 | = | 28000.0 | nM | PMID[485907] |
| NPT526 | Individual Protein | Alpha-glucosidase MAL62 | Saccharomyces cerevisiae | logIC50 | = | 1.602 | n.a. | PMID[485908] |
| NPT1380 | Individual Protein | Avian myoblastosis virus polyprotein II | Avian myeloblastosis virus | Inhibition | = | 40.5 | % | PMID[485909] |
| NPT105 | Individual Protein | Muscleblind-like protein 1 | Homo sapiens | Potency | n.a. | 22387.2 | nM | PMID[485888] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | n.a. | 14125.4 | nM | PMID[485888] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | n.a. | 3981.1 | nM | PMID[485887] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | n.a. | 4466.8 | nM | PMID[485885] |
| NPT49 | Individual Protein | DNA-(apurinic or apyrimidinic site) lyase | Homo sapiens | Potency | n.a. | 5011.9 | nM | PMID[485886] |
| NPT1098 | Individual Protein | Eyes absent homolog 2 | Homo sapiens | Potency | n.a. | 8493.6 | nM | PMID[485894] |
| NPT1120 | Individual Protein | Pancreatic triacylglycerol lipase | Sus scrofa | IC50 | > | 100000.0 | nM | PMID[485914] |
| NPT1120 | Individual Protein | Pancreatic triacylglycerol lipase | Sus scrofa | Inhibition | = | 26.3 | % | PMID[485914] |
| NPT1721 | Individual Protein | Death-associated protein kinase 1 | Homo sapiens | IC50 | = | 1600.0 | nM | PMID[485915] |
| NPT1721 | Individual Protein | Death-associated protein kinase 1 | Homo sapiens | IC50 | = | 11000.0 | nM | PMID[485915] |
| NPT2715 | Individual Protein | Receptor-type tyrosine-protein phosphatase S | Homo sapiens | Inhibition | = | 84.9 | % | PMID[485917] |
| NPT2715 | Individual Protein | Receptor-type tyrosine-protein phosphatase S | Homo sapiens | IC50 | = | 4300.0 | nM | PMID[485917] |
| NPT1443 | Individual Protein | Pyruvate dehydrogenase kinase isoform 1 | Homo sapiens | Inhibition | = | 4.36 | % | PMID[485919] |
| NPT1821 | Individual Protein | Sortase | Staphylococcus aureus | IC50 | = | 19000.0 | nM | PMID[485926] |
| NPT1821 | Individual Protein | Sortase | Staphylococcus aureus | IC50 | = | 37390.0 | nM | PMID[485926] |
| NPT1663 | Individual Protein | Tyrosine-protein kinase YES | Homo sapiens | Ki | = | 7900.0 | nM | PMID[485930] |
| NPT1385 | Individual Protein | Histone-lysine N-methyltransferase SETD7 | Homo sapiens | Inhibition | = | 87.0 | % | PMID[485931] |
| NPT1385 | Individual Protein | Histone-lysine N-methyltransferase SETD7 | Homo sapiens | Inhibition | = | 62.0 | % | PMID[485931] |
| NPT1385 | Individual Protein | Histone-lysine N-methyltransferase SETD7 | Homo sapiens | Inhibition | = | 40.0 | % | PMID[485931] |
| NPT1385 | Individual Protein | Histone-lysine N-methyltransferase SETD7 | Homo sapiens | IC50 | = | 6020.0 | nM | PMID[485931] |
| NPT2266 | Individual Protein | Fructose-1,6-bisphosphatase | Homo sapiens | Inhibition | < | 20.0 | % | PMID[485932] |
| NPT1670 | Individual Protein | DNA-3-methyladenine glycosylase | Homo sapiens | IC50 | = | 2600.0 | nM | PMID[485936] |
| NPT172 | Organism | Proteus mirabilis | Proteus mirabilis | MIC90 | = | 120.0 | ug.mL-1 | PMID[485856] |
| NPT19 | Organism | Escherichia coli | Escherichia coli | MIC90 | = | 1380.0 | ug.mL-1 | PMID[485856] |
| NPT16 | Organism | Staphylococcus aureus | Staphylococcus aureus | MIC90 | = | 120.0 | ug.mL-1 | PMID[485856] |
| NPT18 | Organism | Pseudomonas aeruginosa | Pseudomonas aeruginosa | MIC90 | = | 120.0 | ug.mL-1 | PMID[485856] |
| NPT2554 | Protein Family | Adenosine A2 receptor | Rattus norvegicus | Ki | = | 17300.0 | nM | PMID[485857] |
| NPT2555 | Others | Adenosine receptors; A1 & A3 | Rattus norvegicus | Ratio | = | 0.1 | n.a. | PMID[485857] |
| NPT27 | Others | Unspecified | Ki | = | 30900.0 | nM | PMID[485860] | |
| NPT6 | Organism | Plasmodium falciparum | Plasmodium falciparum | IC50 | = | 110400.0 | nM | PMID[485861] |
| NPT6 | Organism | Plasmodium falciparum | Plasmodium falciparum | IC50 | = | 16100.0 | nM | PMID[485861] |
| NPT24 | Organism | Human immunodeficiency virus 1 | Human immunodeficiency virus 1 | EC50 | < | 100000.0 | nM | PMID[485862] |
| NPT2 | Others | Unspecified | Activity | = | 18.0 | % | PMID[485865] | |
| NPT2 | Others | Unspecified | Activity | = | 31.5 | % | PMID[485865] | |
| NPT2 | Others | Unspecified | Activity | = | 13.3 | % | PMID[485865] | |
| NPT2 | Others | Unspecified | IC50 | = | 430.0 | nM | PMID[485865] | |
| NPT2 | Others | Unspecified | IC50 | = | 730.0 | nM | PMID[485865] | |
| NPT2 | Others | Unspecified | IC50 | = | 120.0 | nM | PMID[485865] | |
| NPT566 | Organism | Salmonella typhimurium | Salmonella enterica subsp. enterica serovar Typhimurium | Inhibition | < | 30.0 | % | PMID[485866] |
| NPT566 | Organism | Salmonella typhimurium | Salmonella enterica subsp. enterica serovar Typhimurium | Inhibition | < | 20.0 | % | PMID[485866] |
| NPT566 | Organism | Salmonella typhimurium | Salmonella enterica subsp. enterica serovar Typhimurium | Inhibition | = | 0.0 | % | PMID[485866] |
| NPT35 | Others | n.a. | Activity | = | 0.83 | mmol/L | PMID[485868] | |
| NPT1084 | Subcellular | Liver microsomes | Rattus norvegicus | IC50 | = | 23000000.0 | nM | PMID[485868] |
| NPT2 | Others | Unspecified | ID50 | = | 1.03 | uM | PMID[485869] | |
| NPT2 | Others | Unspecified | ID50 | = | 1.01 | uM | PMID[485869] | |
| NPT2 | Others | Unspecified | FC | = | 10.0 | n.a. | PMID[485869] | |
| NPT967 | Individual Protein | Xanthine dehydrogenase | Homo sapiens | IC50 | = | 10100.0 | nM | PMID[485872] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | IC50 | = | 9100.0 | nM | PMID[485872] |
| NPT2 | Others | Unspecified | Activity | = | 0.0 | n.a. | PMID[485873] | |
| NPT2 | Others | Unspecified | IC50 | = | 110800.0 | nM | PMID[485876] | |
| NPT24 | Organism | Human immunodeficiency virus 1 | Human immunodeficiency virus 1 | EC50 | > | 331000.0 | nM | PMID[485877] |
| NPT2 | Others | Unspecified | IC50 | = | 50.0 | ug.mL-1 | PMID[485878] | |
| NPT2 | Others | Unspecified | IC50 | = | 42.1 | ug.mL-1 | PMID[485875] | |
| NPT831 | Individual Protein | Serine/threonine-protein kinase PLK1 | Homo sapiens | IC50 | = | 12600.0 | nM | PMID[485879] |
| NPT35 | Others | n.a. | Activity | = | 165.3 | mg/L | PMID[485880] | |
| NPT35 | Others | n.a. | Activity | = | 2.68 | MU | PMID[485880] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Activity | = | 2.55 | MU | PMID[485880] |
| NPT35 | Others | n.a. | Activity | = | 2.75 | MU | PMID[485880] | |
| NPT2 | Others | Unspecified | Ratio IC50 | = | 20.0 | n.a. | PMID[485881] | |
| NPT2 | Others | Unspecified | Inhibition | = | 94.0 | % | PMID[485881] | |
| NPT2 | Others | Unspecified | IC50 | = | 670.0 | nM | PMID[485881] | |
| NPT2 | Others | Unspecified | IC50 | = | 240.0 | nM | PMID[485881] | |
| NPT2 | Others | Unspecified | IC50 | = | 49500.0 | nM | PMID[485881] | |
| NPT2 | Others | Unspecified | Inhibition | = | 99.0 | % | PMID[485881] | |
| NPT2 | Others | Unspecified | Ratio IC50 | = | 10.0 | n.a. | PMID[485881] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Activity | = | 81.5 | % | PMID[485882] |
| NPT566 | Organism | Salmonella typhimurium | Salmonella enterica subsp. enterica serovar Typhimurium | pID50 | = | 6.155 | n.a. | PMID[485883] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 28183.8 | nM | PMID[485885] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 22387.2 | nM | PMID[485886] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 35481.3 | nM | PMID[485887] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 22387.2 | nM | PMID[485888] |
| NPT2 | Others | Unspecified | Potency | = | 28183.8 | nM | PMID[485885] | |
| NPT2 | Others | Unspecified | Potency | = | 39810.7 | nM | PMID[485888] | |
| NPT312 | Organism | Saccharomyces cerevisiae | Saccharomyces cerevisiae | Potency | = | 18356.4 | nM | PMID[485890] |
| NPT2 | Others | Unspecified | Potency | = | 12589.3 | nM | PMID[485885] | |
| NPT794 | Individual Protein | Endonuclease 4 | Escherichia coli K-12 | Potency | = | 3548.1 | nM | PMID[485885] |
| NPT94 | Individual Protein | Aldehyde dehydrogenase 1A1 | Homo sapiens | Potency | = | 19952.6 | nM | PMID[485886] |
| NPT792 | Individual Protein | Arachidonate 15-lipoxygenase | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485888] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 11220.2 | nM | PMID[485887] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485885] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 15848.9 | nM | PMID[485886] |
| NPT197 | Protein-Protein Interaction | Menin/Histone-lysine N-methyltransferase MLL | Homo sapiens | Potency | = | 19952.6 | nM | PMID[485888] |
| NPT95 | Individual Protein | Muscarinic acetylcholine receptor M1 | Rattus norvegicus | Potency | = | 20.0 | nM | PMID[485888] |
| NPT96 | Individual Protein | Thrombopoietin | Homo sapiens | Potency | = | 3162.3 | nM | PMID[485888] |
| NPT800 | Individual Protein | M-phase phosphoprotein 8 | Homo sapiens | Potency | n.a. | 39810.7 | nM | PMID[485889] |
| NPT800 | Individual Protein | M-phase phosphoprotein 8 | Homo sapiens | Potency | n.a. | 21192.3 | nM | PMID[485890] |
| NPT2 | Others | Unspecified | Potency | n.a. | 3662.6 | nM | PMID[485888] | |
| NPT755 | Individual Protein | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | Homo sapiens | Potency | n.a. | 3793.3 | nM | PMID[485890] |
| NPT533 | Protein-Protein Interaction | Runt-related transcription factor 1/Core-binding factor subunit beta | Homo sapiens | Potency | n.a. | 14125.4 | nM | PMID[485887] |
| NPT533 | Protein-Protein Interaction | Runt-related transcription factor 1/Core-binding factor subunit beta | Homo sapiens | Potency | n.a. | 28183.8 | nM | PMID[485888] |
| NPT533 | Protein-Protein Interaction | Runt-related transcription factor 1/Core-binding factor subunit beta | Homo sapiens | Potency | n.a. | 25118.9 | nM | PMID[485886] |
| NPT803 | Individual Protein | Flap endonuclease 1 | Homo sapiens | Potency | n.a. | 2377.8 | nM | PMID[485890] |
| NPT740 | Individual Protein | Beta-secretase 1 | Homo sapiens | IC50 | = | 12300.0 | nM | PMID[485893] |
| NPT7 | Individual Protein | Thioredoxin reductase 1, cytoplasmic | Rattus norvegicus | Potency | n.a. | 7079.5 | nM | PMID[485894] |
| NPT7 | Individual Protein | Thioredoxin reductase 1, cytoplasmic | Rattus norvegicus | Potency | n.a. | 29934.9 | nM | PMID[485890] |
| NPT7 | Individual Protein | Thioredoxin reductase 1, cytoplasmic | Rattus norvegicus | Potency | n.a. | 79432.8 | nM | PMID[485889] |
| NPT803 | Individual Protein | Flap endonuclease 1 | Homo sapiens | Potency | n.a. | 28183.8 | nM | PMID[485894] |
| NPT698 | Individual Protein | Regulator of G-protein signaling 4 | Homo sapiens | Potency | n.a. | 29934.9 | nM | PMID[485890] |
| NPT8 | Individual Protein | DNA polymerase iota | Homo sapiens | Potency | n.a. | 50118.7 | nM | PMID[485894] |
| NPT8 | Individual Protein | DNA polymerase iota | Homo sapiens | Potency | n.a. | 3981.1 | nM | PMID[485885] |
| NPT9 | Individual Protein | DNA polymerase eta | Homo sapiens | Potency | n.a. | 7943.3 | nM | PMID[485885] |
| NPT98 | Individual Protein | HERG | Homo sapiens | Potency | n.a. | 3162.3 | nM | PMID[485885] |
| NPT2 | Others | Unspecified | Potency | n.a. | 49.9 | nM | PMID[485890] | |
| NPT191 | Organism | Hepatitis C virus | Hepatitis C virus | Inhibition | = | 31.09 | % | PMID[485899] |
| NPT27 | Others | Unspecified | CC50 | > | 50000.0 | nM | PMID[485899] | |
| NPT2 | Others | Unspecified | IC50 | = | 26000.0 | nM | PMID[485904] | |
| NPT2 | Others | Unspecified | IC50 | = | 2300.0 | nM | PMID[485904] | |
| NPT72 | Individual Protein | Solute carrier organic anion transporter family member 1B3 | Homo sapiens | Inhibition | = | 78.5 | % | PMID[485905] |
| NPT73 | Individual Protein | Solute carrier organic anion transporter family member 1B1 | Homo sapiens | Inhibition | = | 85.2 | % | PMID[485905] |
| NPT67 | Individual Protein | Cholinesterase | Equus caballus | Inhibition | = | -3.98 | % | PMID[485906] |
| NPT1 | Others | Radical scavenging activity | Inhibition | = | 96.5 | % | PMID[485910] | |
| NPT740 | Individual Protein | Beta-secretase 1 | Homo sapiens | IC50 | = | 21700.0 | nM | PMID[485911] |
| NPT518 | Protein Complex | Tubulin | Homo sapiens | Kd | = | 199526.23 | nM | PMID[485912] |
| NPT518 | Protein Complex | Tubulin | Homo sapiens | Kd | > | 100000.0 | nM | PMID[485912] |
| NPT72 | Individual Protein | Solute carrier organic anion transporter family member 1B3 | Homo sapiens | Inhibition | = | 45.37 | % | PMID[485913] |
| NPT73 | Individual Protein | Solute carrier organic anion transporter family member 1B1 | Homo sapiens | Inhibition | = | 24.17 | % | PMID[485913] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Potency | n.a. | 21117.7 | nM | PMID[485891] |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Potency | n.a. | 10583.9 | nM | PMID[485891] |
| NPT2 | Others | Unspecified | Potency | n.a. | 3.1 | nM | PMID[485890] | |
| NPT2 | Others | Unspecified | Potency | n.a. | 16946.9 | nM | PMID[485894] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Potency | n.a. | 7.5 | nM | PMID[485891] |
| NPT2 | Others | Unspecified | Potency | n.a. | 10000.0 | nM | PMID[485890] | |
| NPT20529 | NON-MOLECULAR | NON-PROTEIN TARGET | n.a. | Potency | n.a. | 9432.9 | nM | PMID[485891] |
| NPT1609 | Protein Complex | Casein kinase II | Homo sapiens | IC50 | = | 10000.0 | nM | PMID[485915] |
| NPT78 | Individual Protein | Beta amyloid A4 protein | Homo sapiens | IC50 | = | 30300.0 | nM | PMID[485918] |
| NPT35 | Others | n.a. | Stability | > | 80.0 | % | PMID[485918] | |
| NPT78 | Individual Protein | Beta amyloid A4 protein | Homo sapiens | TIME | = | 10.0 | hr | PMID[485918] |
| NPT2 | Others | Unspecified | IC50 | = | 30300.0 | nM | PMID[485920] | |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | Inhibition | = | -4.08 | % | PMID[485921] |
| NPT22879 | SINGLE PROTEIN | DNA ligase 1 | Homo sapiens | IC50 | = | 68.0 | ug.mL-1 | PMID[485922] |
| NPT21765 | SINGLE PROTEIN | Sortase A | Streptococcus mutans | IC50 | = | 27200.0 | nM | PMID[485923] |
| NPT22915 | PROTEIN FAMILY | Casein kinase 2 | Homo sapiens | IC50 | = | 10000.0 | nM | PMID[485924] |
| NPT21773 | SINGLE PROTEIN | Thiosulfate sulfurtransferase | Homo sapiens | IC50 | = | 57000.0 | nM | PMID[485925] |
| NPT2 | Others | Unspecified | IC50 | = | 47000.0 | nM | PMID[485925] | |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | IC50 | = | 5200.0 | nM | PMID[485925] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | IC50 | = | 8500.0 | nM | PMID[485925] |
| NPT21774 | PROTEIN COMPLEX | HSP60/HSP10 | Homo sapiens | IC50 | = | 11000.0 | nM | PMID[485925] |
| NPT21775 | SINGLE PROTEIN | 60 kDa chaperonin | Escherichia coli (strain K12) | IC50 | > | 250000.0 | nM | PMID[485925] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | Inhibition | = | 98.0 | % | PMID[485925] |
| NPT21772 | PROTEIN COMPLEX | GroEL/GroES | Escherichia coli | Inhibition | = | 97.0 | % | PMID[485925] |
| NPT2 | Others | Unspecified | IC50 | = | 70000.0 | nM | PMID[485927] | |
| NPT2 | Others | Unspecified | Inhibition | = | 100.0 | % | PMID[485927] | |
| NPT20556 | SINGLE PROTEIN | Replicase polyprotein 1ab | Severe acute respiratory syndrome coronavirus 2 | Inhibition | = | 9.068 | % | PMID[485928] |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | Inhibition | = | -0.19 | % | PMID[485929] |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | IC50 | > | 20000.0 | nM | PMID[485933] |
| NPT20555 | ORGANISM | SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 | IC50 | > | 19952.62 | nM | PMID[485933] |
| NPT78 | Individual Protein | Beta amyloid A4 protein | Homo sapiens | IC50 | = | 8500.0 | nM | PMID[485934] |
| NPT2 | Others | Unspecified | IC50 | = | 9000.0 | nM | PMID[485935] |
☑ Note for Activity Records:
☉ The quantitative biological activities were primarily integrated from ChEMBL (Version-30) database and were also directly collected from PubMed literature. PubMed PMID was provided as the reference link for each activity record.
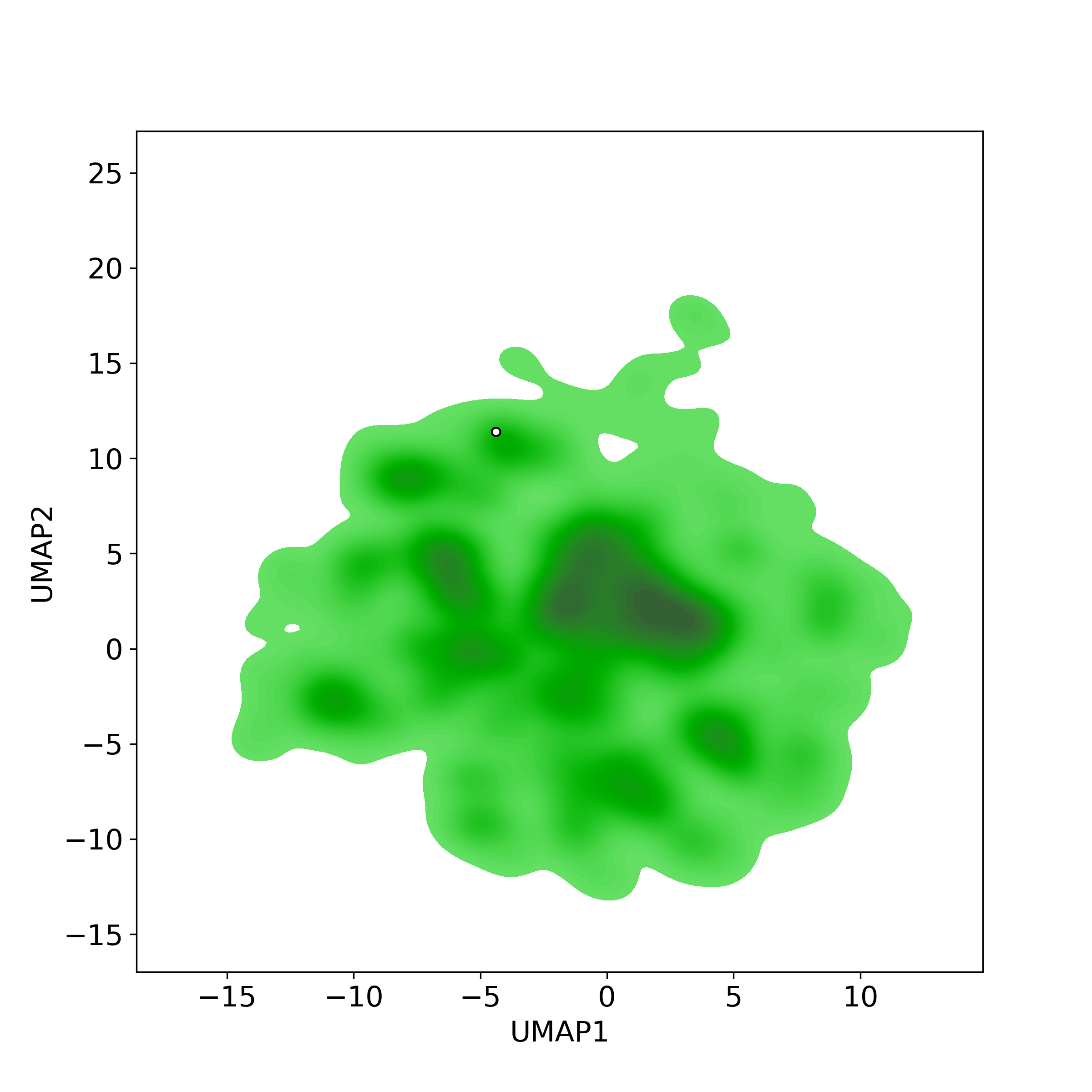
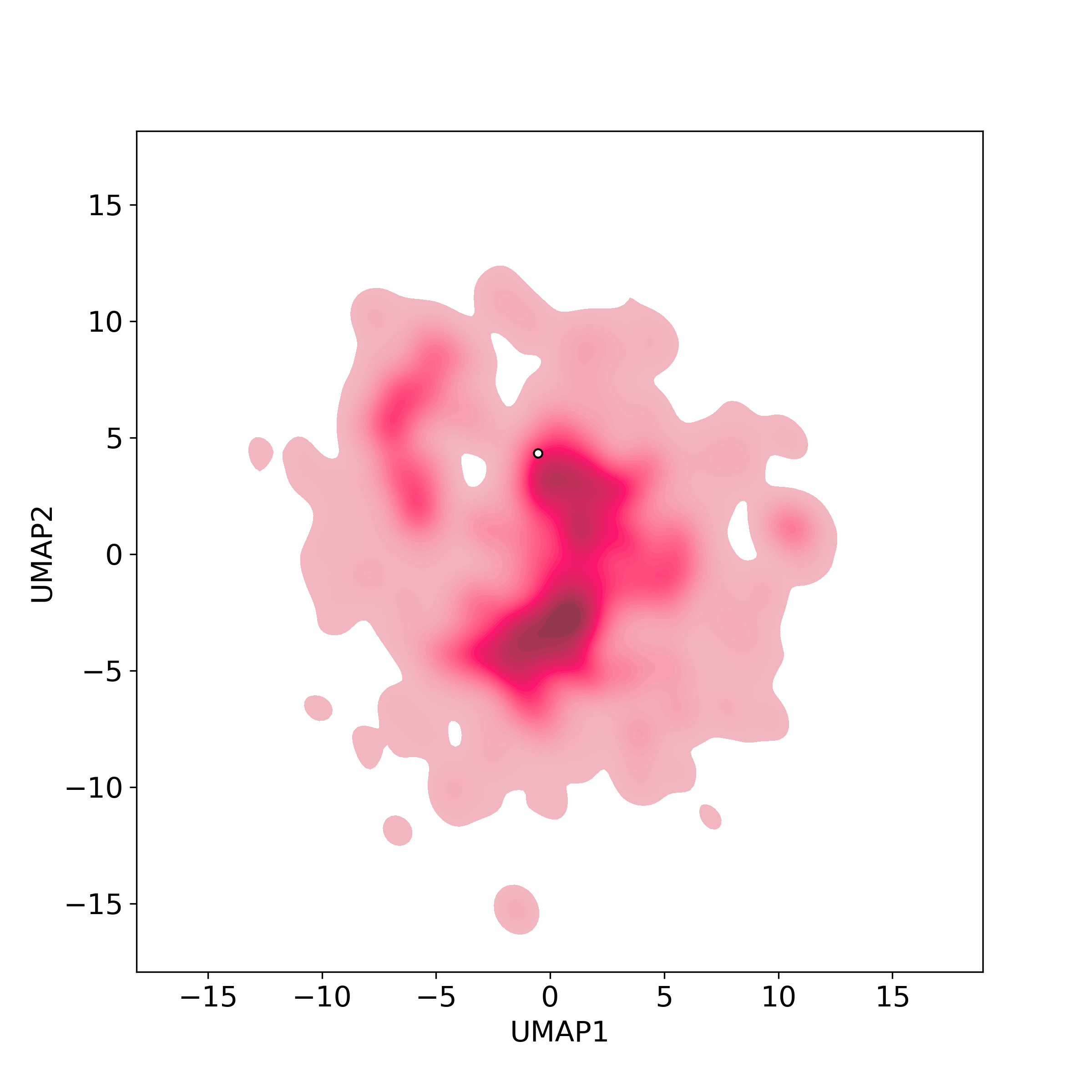
 Chemically structural similarity: I. Similar Active Natural Products in NPASS
Chemically structural similarity: I. Similar Active Natural Products in NPASSTop-200 similar NPs were calculated against the active-NP-set (includes 4,3285 NPs with experimentally-derived bioactivity available in NPASS)
Similarity level is defined by Tanimoto coefficient (Tc) between two molecules. Tc lies between [0, 1] where '1' indicates the highest similarity. What is Tanimoto coefficient 
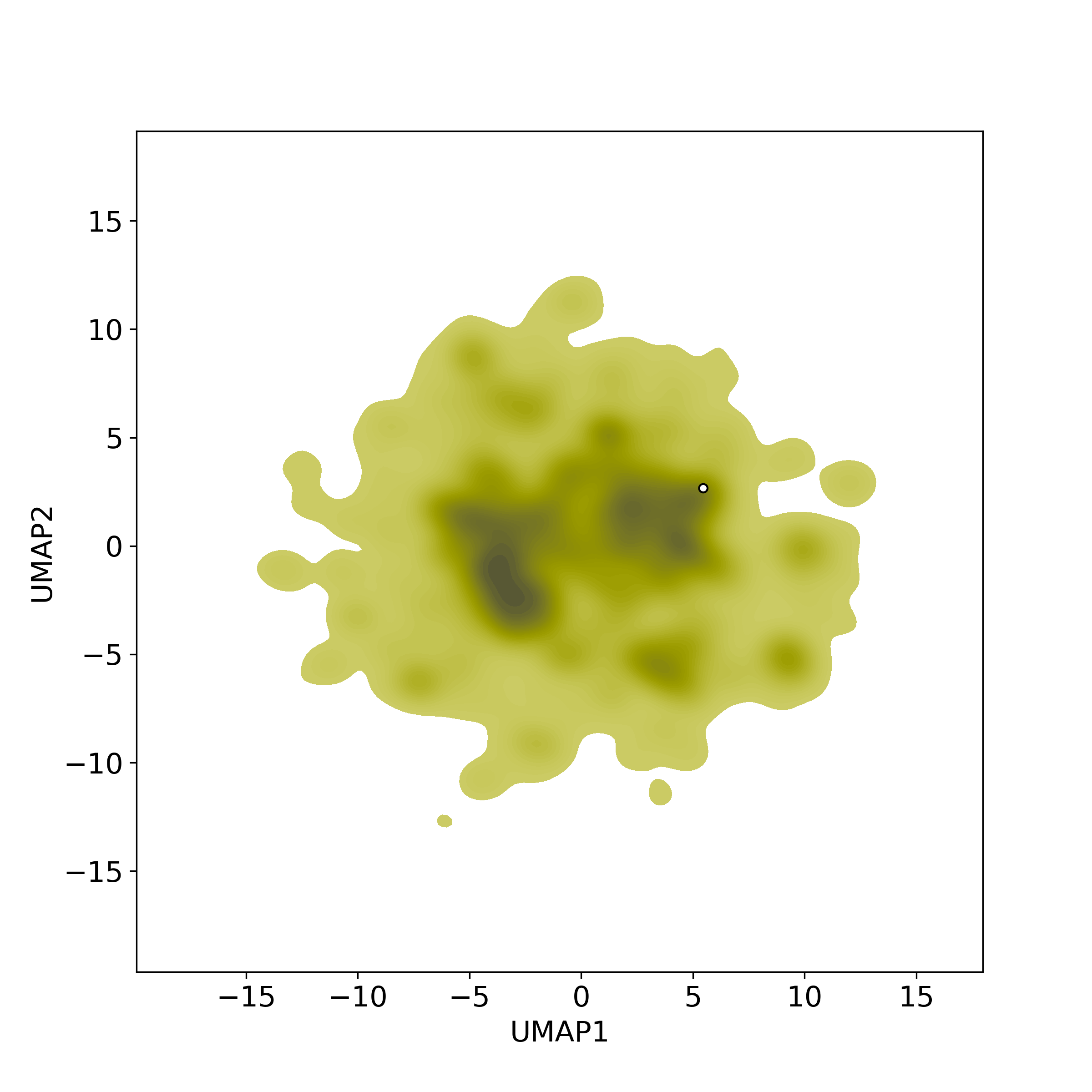
● The left chart: Distribution of similarity level between NPC216361 and all remaining natural products in the NPASS database.
● The right table: Most similar natural products (Tc>=0.56 or Top200).
| Similarity Score | Similarity Level | Natural Product ID |
|---|---|---|
| 1.0 | High Similarity | NPC281917 |
| 1.0 | High Similarity | NPC187432 |
| 1.0 | High Similarity | NPC256042 |
| 1.0 | High Similarity | NPC116775 |
| 0.9924 | High Similarity | NPC299379 |
| 0.9846 | High Similarity | NPC248872 |
| 0.9774 | High Similarity | NPC152042 |
| 0.9774 | High Similarity | NPC143799 |
| 0.9774 | High Similarity | NPC241838 |
| 0.9769 | High Similarity | NPC274121 |
| 0.9769 | High Similarity | NPC50898 |
| 0.9769 | High Similarity | NPC78540 |
| 0.9769 | High Similarity | NPC213216 |
| 0.9701 | High Similarity | NPC184536 |
| 0.9701 | High Similarity | NPC146679 |
| 0.9701 | High Similarity | NPC59951 |
| 0.9701 | High Similarity | NPC269652 |
| 0.9701 | High Similarity | NPC281207 |
| 0.9701 | High Similarity | NPC103342 |
| 0.9701 | High Similarity | NPC103904 |
| 0.9701 | High Similarity | NPC230285 |
| 0.9695 | High Similarity | NPC172262 |
| 0.9692 | High Similarity | NPC128216 |
| 0.963 | High Similarity | NPC262094 |
| 0.963 | High Similarity | NPC136840 |
| 0.963 | High Similarity | NPC90582 |
| 0.9618 | High Similarity | NPC57601 |
| 0.9615 | High Similarity | NPC113006 |
| 0.9559 | High Similarity | NPC200694 |
| 0.9559 | High Similarity | NPC473042 |
| 0.9552 | High Similarity | NPC276905 |
| 0.9549 | High Similarity | NPC473887 |
| 0.9549 | High Similarity | NPC231772 |
| 0.9549 | High Similarity | NPC194281 |
| 0.9549 | High Similarity | NPC47815 |
| 0.9549 | High Similarity | NPC234133 |
| 0.9549 | High Similarity | NPC13408 |
| 0.9549 | High Similarity | NPC29353 |
| 0.9549 | High Similarity | NPC124784 |
| 0.9549 | High Similarity | NPC127447 |
| 0.9489 | High Similarity | NPC293053 |
| 0.9489 | High Similarity | NPC24821 |
| 0.9489 | High Similarity | NPC9117 |
| 0.9489 | High Similarity | NPC470216 |
| 0.9489 | High Similarity | NPC212932 |
| 0.9489 | High Similarity | NPC190637 |
| 0.9478 | High Similarity | NPC254841 |
| 0.9478 | High Similarity | NPC250266 |
| 0.9478 | High Similarity | NPC266597 |
| 0.9474 | High Similarity | NPC239495 |
| 0.9474 | High Similarity | NPC9985 |
| 0.942 | High Similarity | NPC216538 |
| 0.942 | High Similarity | NPC273538 |
| 0.942 | High Similarity | NPC326500 |
| 0.9407 | High Similarity | NPC201395 |
| 0.9389 | High Similarity | NPC137264 |
| 0.9389 | High Similarity | NPC5515 |
| 0.9389 | High Similarity | NPC270369 |
| 0.9385 | High Similarity | NPC197425 |
| 0.9353 | High Similarity | NPC153758 |
| 0.9353 | High Similarity | NPC178343 |
| 0.9353 | High Similarity | NPC179271 |
| 0.9353 | High Similarity | NPC306488 |
| 0.9353 | High Similarity | NPC20791 |
| 0.9353 | High Similarity | NPC5820 |
| 0.9353 | High Similarity | NPC124729 |
| 0.9348 | High Similarity | NPC158874 |
| 0.9343 | High Similarity | NPC310128 |
| 0.9338 | High Similarity | NPC301217 |
| 0.9338 | High Similarity | NPC96565 |
| 0.9338 | High Similarity | NPC303633 |
| 0.9338 | High Similarity | NPC216978 |
| 0.9338 | High Similarity | NPC55018 |
| 0.9338 | High Similarity | NPC18260 |
| 0.9338 | High Similarity | NPC220062 |
| 0.9338 | High Similarity | NPC78913 |
| 0.9333 | High Similarity | NPC296490 |
| 0.9333 | High Similarity | NPC275055 |
| 0.9333 | High Similarity | NPC295261 |
| 0.9333 | High Similarity | NPC13768 |
| 0.9333 | High Similarity | NPC79943 |
| 0.9333 | High Similarity | NPC107586 |
| 0.9333 | High Similarity | NPC287246 |
| 0.9333 | High Similarity | NPC12296 |
| 0.9333 | High Similarity | NPC32441 |
| 0.9333 | High Similarity | NPC290291 |
| 0.9333 | High Similarity | NPC243083 |
| 0.9333 | High Similarity | NPC228661 |
| 0.9318 | High Similarity | NPC475589 |
| 0.9318 | High Similarity | NPC31872 |
| 0.9318 | High Similarity | NPC473584 |
| 0.9313 | High Similarity | NPC39753 |
| 0.9313 | High Similarity | NPC115998 |
| 0.9286 | High Similarity | NPC74881 |
| 0.9286 | High Similarity | NPC51443 |
| 0.9286 | High Similarity | NPC14001 |
| 0.9286 | High Similarity | NPC166757 |
| 0.9281 | High Similarity | NPC62290 |
| 0.9281 | High Similarity | NPC4152 |
| 0.9281 | High Similarity | NPC326506 |
| 0.9281 | High Similarity | NPC49130 |
| 0.9281 | High Similarity | NPC279417 |
| 0.9281 | High Similarity | NPC208176 |
| 0.9281 | High Similarity | NPC306607 |
| 0.9281 | High Similarity | NPC142731 |
| 0.9275 | High Similarity | NPC471520 |
| 0.927 | High Similarity | NPC270883 |
| 0.927 | High Similarity | NPC159275 |
| 0.927 | High Similarity | NPC241100 |
| 0.927 | High Similarity | NPC261227 |
| 0.927 | High Similarity | NPC172986 |
| 0.9265 | High Similarity | NPC99333 |
| 0.9265 | High Similarity | NPC472460 |
| 0.9265 | High Similarity | NPC188947 |
| 0.9265 | High Similarity | NPC147686 |
| 0.9265 | High Similarity | NPC280284 |
| 0.9265 | High Similarity | NPC329225 |
| 0.9265 | High Similarity | NPC118813 |
| 0.9259 | High Similarity | NPC61546 |
| 0.9259 | High Similarity | NPC72452 |
| 0.9248 | High Similarity | NPC25937 |
| 0.9248 | High Similarity | NPC10971 |
| 0.922 | High Similarity | NPC296869 |
| 0.922 | High Similarity | NPC244407 |
| 0.9214 | High Similarity | NPC229190 |
| 0.9214 | High Similarity | NPC220418 |
| 0.9214 | High Similarity | NPC169749 |
| 0.9214 | High Similarity | NPC470458 |
| 0.9214 | High Similarity | NPC148011 |
| 0.9214 | High Similarity | NPC1940 |
| 0.9209 | High Similarity | NPC279121 |
| 0.9203 | High Similarity | NPC118840 |
| 0.9203 | High Similarity | NPC110969 |
| 0.9203 | High Similarity | NPC147688 |
| 0.9203 | High Similarity | NPC64908 |
| 0.9203 | High Similarity | NPC282300 |
| 0.9203 | High Similarity | NPC156590 |
| 0.9203 | High Similarity | NPC14871 |
| 0.9203 | High Similarity | NPC205006 |
| 0.9197 | High Similarity | NPC329203 |
| 0.9197 | High Similarity | NPC150648 |
| 0.9197 | High Similarity | NPC261234 |
| 0.9197 | High Similarity | NPC53181 |
| 0.9197 | High Similarity | NPC217186 |
| 0.9197 | High Similarity | NPC225153 |
| 0.9197 | High Similarity | NPC20709 |
| 0.9197 | High Similarity | NPC316480 |
| 0.9197 | High Similarity | NPC274784 |
| 0.9197 | High Similarity | NPC310135 |
| 0.9197 | High Similarity | NPC265871 |
| 0.9197 | High Similarity | NPC222342 |
| 0.9191 | High Similarity | NPC84585 |
| 0.9191 | High Similarity | NPC188879 |
| 0.9191 | High Similarity | NPC160972 |
| 0.9191 | High Similarity | NPC476480 |
| 0.9185 | High Similarity | NPC234560 |
| 0.9185 | High Similarity | NPC39426 |
| 0.9179 | High Similarity | NPC223457 |
| 0.9179 | High Similarity | NPC93756 |
| 0.9179 | High Similarity | NPC108113 |
| 0.9173 | High Similarity | NPC278556 |
| 0.9173 | High Similarity | NPC65060 |
| 0.9155 | High Similarity | NPC195351 |
| 0.9155 | High Similarity | NPC270465 |
| 0.9155 | High Similarity | NPC269420 |
| 0.9155 | High Similarity | NPC470461 |
| 0.9155 | High Similarity | NPC159103 |
| 0.9155 | High Similarity | NPC87125 |
| 0.9149 | High Similarity | NPC211466 |
| 0.9149 | High Similarity | NPC40086 |
| 0.9149 | High Similarity | NPC3779 |
| 0.9149 | High Similarity | NPC472918 |
| 0.9149 | High Similarity | NPC476182 |
| 0.9149 | High Similarity | NPC44721 |
| 0.9149 | High Similarity | NPC259632 |
| 0.9149 | High Similarity | NPC122828 |
| 0.9149 | High Similarity | NPC176869 |
| 0.9143 | High Similarity | NPC71210 |
| 0.9143 | High Similarity | NPC70136 |
| 0.9137 | High Similarity | NPC62840 |
| 0.9137 | High Similarity | NPC214236 |
| 0.9137 | High Similarity | NPC59739 |
| 0.9137 | High Similarity | NPC11561 |
| 0.9137 | High Similarity | NPC226636 |
| 0.9137 | High Similarity | NPC159855 |
| 0.9137 | High Similarity | NPC169479 |
| 0.9137 | High Similarity | NPC217083 |
| 0.9137 | High Similarity | NPC175013 |
| 0.9137 | High Similarity | NPC299080 |
| 0.9137 | High Similarity | NPC293852 |
| 0.9137 | High Similarity | NPC78803 |
| 0.913 | High Similarity | NPC235239 |
| 0.913 | High Similarity | NPC129853 |
| 0.913 | High Similarity | NPC110228 |
| 0.913 | High Similarity | NPC69769 |
| 0.913 | High Similarity | NPC6407 |
| 0.913 | High Similarity | NPC475680 |
| 0.913 | High Similarity | NPC188243 |
| 0.913 | High Similarity | NPC150522 |
| 0.913 | High Similarity | NPC76445 |
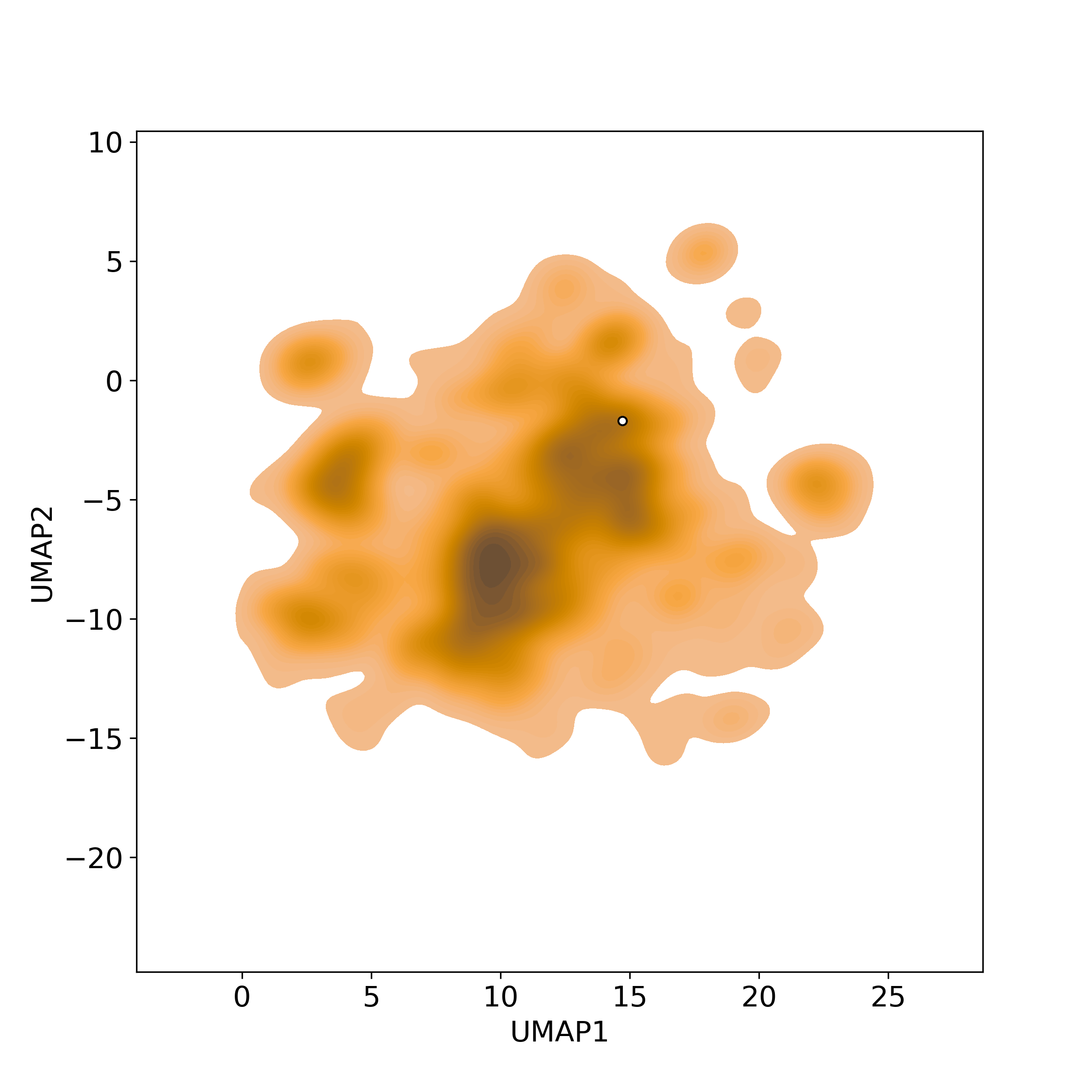
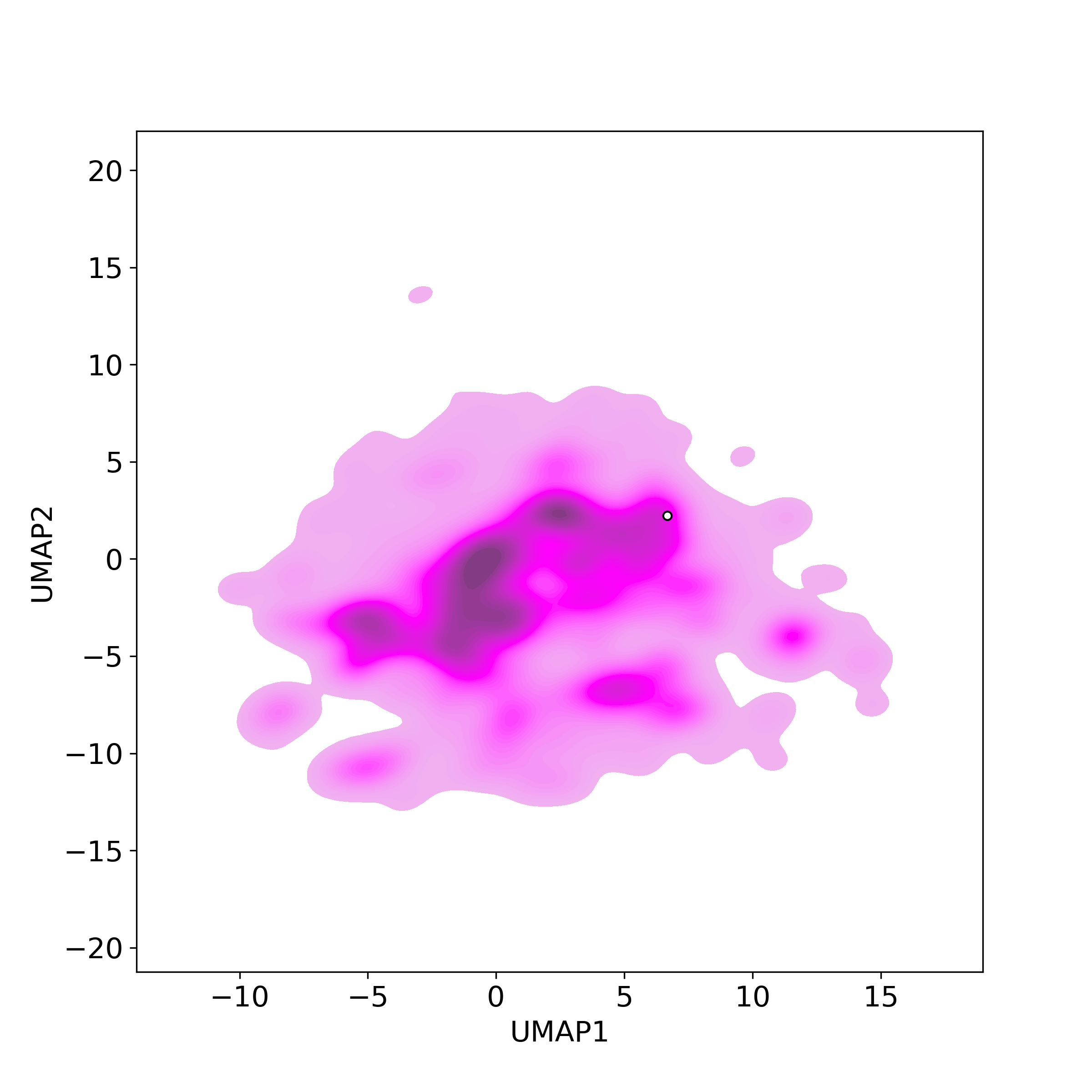
 Chemically structural similarity: II. Similar Clinical/Approved Drugs
Chemically structural similarity: II. Similar Clinical/Approved DrugsSimilarity level is defined by Tanimoto coefficient (Tc) between two molecules.
● The left chart: Distribution of similarity level between NPC216361 and all drugs/candidates.
● The right table: Most similar clinical/approved drugs (Tc>=0.56 or Top200).
| Similarity Score | Similarity Level | Drug ID | Developmental Stage |
|---|---|---|---|
| 0.942 | High Similarity | NPD4378 | Clinical (unspecified phase) |
| 0.9353 | High Similarity | NPD1512 | Approved |
| 0.9333 | High Similarity | NPD1552 | Clinical (unspecified phase) |
| 0.9333 | High Similarity | NPD1550 | Clinical (unspecified phase) |
| 0.9265 | High Similarity | NPD1549 | Phase 2 |
| 0.9209 | High Similarity | NPD1511 | Approved |
| 0.9185 | High Similarity | NPD1510 | Phase 2 |
| 0.903 | High Similarity | NPD1240 | Approved |
| 0.8897 | High Similarity | NPD1607 | Approved |
| 0.8873 | High Similarity | NPD7410 | Clinical (unspecified phase) |
| 0.8705 | High Similarity | NPD2796 | Approved |
| 0.8671 | High Similarity | NPD6799 | Approved |
| 0.8649 | High Similarity | NPD2801 | Approved |
| 0.8592 | High Similarity | NPD3750 | Approved |
| 0.8581 | High Similarity | NPD1934 | Approved |
| 0.8571 | High Similarity | NPD4380 | Phase 2 |
| 0.8523 | High Similarity | NPD2393 | Clinical (unspecified phase) |
| 0.8477 | Intermediate Similarity | NPD7075 | Discontinued |
| 0.8477 | Intermediate Similarity | NPD4381 | Clinical (unspecified phase) |
| 0.8467 | Intermediate Similarity | NPD3817 | Phase 2 |
| 0.8456 | Intermediate Similarity | NPD6801 | Discontinued |
| 0.8451 | Intermediate Similarity | NPD970 | Clinical (unspecified phase) |
| 0.844 | Intermediate Similarity | NPD1551 | Phase 2 |
| 0.8411 | Intermediate Similarity | NPD3882 | Suspended |
| 0.84 | Intermediate Similarity | NPD7096 | Clinical (unspecified phase) |
| 0.8389 | Intermediate Similarity | NPD7411 | Suspended |
| 0.837 | Intermediate Similarity | NPD1203 | Approved |
| 0.8367 | Intermediate Similarity | NPD5403 | Approved |
| 0.8356 | Intermediate Similarity | NPD5401 | Approved |
| 0.8344 | Intermediate Similarity | NPD8443 | Clinical (unspecified phase) |
| 0.8322 | Intermediate Similarity | NPD6599 | Discontinued |
| 0.8269 | Intermediate Similarity | NPD6168 | Clinical (unspecified phase) |
| 0.8269 | Intermediate Similarity | NPD6167 | Clinical (unspecified phase) |
| 0.8269 | Intermediate Similarity | NPD6166 | Phase 2 |
| 0.8264 | Intermediate Similarity | NPD2800 | Approved |
| 0.8239 | Intermediate Similarity | NPD3748 | Approved |
| 0.8235 | Intermediate Similarity | NPD3749 | Approved |
| 0.8228 | Intermediate Similarity | NPD7804 | Clinical (unspecified phase) |
| 0.8201 | Intermediate Similarity | NPD6859 | Clinical (unspecified phase) |
| 0.8182 | Intermediate Similarity | NPD1548 | Phase 1 |
| 0.8158 | Intermediate Similarity | NPD7819 | Suspended |
| 0.8138 | Intermediate Similarity | NPD1243 | Approved |
| 0.8125 | Intermediate Similarity | NPD6797 | Phase 2 |
| 0.8105 | Intermediate Similarity | NPD5402 | Approved |
| 0.8075 | Intermediate Similarity | NPD7251 | Discontinued |
| 0.8074 | Intermediate Similarity | NPD422 | Phase 1 |
| 0.8071 | Intermediate Similarity | NPD2313 | Discontinued |
| 0.805 | Intermediate Similarity | NPD3818 | Discontinued |
| 0.8025 | Intermediate Similarity | NPD7808 | Phase 3 |
| 0.8025 | Intermediate Similarity | NPD4338 | Clinical (unspecified phase) |
| 0.8015 | Intermediate Similarity | NPD9717 | Approved |
| 0.8014 | Intermediate Similarity | NPD7421 | Clinical (unspecified phase) |
| 0.8 | Intermediate Similarity | NPD7054 | Approved |
| 0.8 | Intermediate Similarity | NPD920 | Approved |
| 0.8 | Intermediate Similarity | NPD2344 | Approved |
| 0.7987 | Intermediate Similarity | NPD642 | Clinical (unspecified phase) |
| 0.7973 | Intermediate Similarity | NPD643 | Clinical (unspecified phase) |
| 0.7962 | Intermediate Similarity | NPD6959 | Discontinued |
| 0.7959 | Intermediate Similarity | NPD1878 | Clinical (unspecified phase) |
| 0.7958 | Intermediate Similarity | NPD943 | Approved |
| 0.795 | Intermediate Similarity | NPD7472 | Approved |
| 0.7941 | Intermediate Similarity | NPD1547 | Clinical (unspecified phase) |
| 0.7935 | Intermediate Similarity | NPD7768 | Phase 2 |
| 0.7925 | Intermediate Similarity | NPD7852 | Clinical (unspecified phase) |
| 0.7899 | Intermediate Similarity | NPD3225 | Approved |
| 0.7847 | Intermediate Similarity | NPD6651 | Approved |
| 0.784 | Intermediate Similarity | NPD7074 | Phase 3 |
| 0.7821 | Intermediate Similarity | NPD4868 | Clinical (unspecified phase) |
| 0.7817 | Intermediate Similarity | NPD3268 | Approved |
| 0.781 | Intermediate Similarity | NPD1610 | Phase 2 |
| 0.7808 | Intermediate Similarity | NPD2935 | Discontinued |
| 0.7785 | Intermediate Similarity | NPD2309 | Approved |
| 0.774 | Intermediate Similarity | NPD651 | Clinical (unspecified phase) |
| 0.774 | Intermediate Similarity | NPD1509 | Clinical (unspecified phase) |
| 0.774 | Intermediate Similarity | NPD2799 | Discontinued |
| 0.7736 | Intermediate Similarity | NPD1247 | Approved |
| 0.7722 | Intermediate Similarity | NPD919 | Approved |
| 0.7714 | Intermediate Similarity | NPD2797 | Approved |
| 0.7697 | Intermediate Similarity | NPD4419 | Clinical (unspecified phase) |
| 0.7692 | Intermediate Similarity | NPD1465 | Phase 2 |
| 0.7692 | Intermediate Similarity | NPD1296 | Phase 2 |
| 0.7688 | Intermediate Similarity | NPD6232 | Discontinued |
| 0.7687 | Intermediate Similarity | NPD6100 | Approved |
| 0.7687 | Intermediate Similarity | NPD6099 | Approved |
| 0.7676 | Intermediate Similarity | NPD6832 | Phase 2 |
| 0.7676 | Intermediate Similarity | NPD1529 | Clinical (unspecified phase) |
| 0.7673 | Intermediate Similarity | NPD5494 | Approved |
| 0.7669 | Intermediate Similarity | NPD7286 | Phase 2 |
| 0.7662 | Intermediate Similarity | NPD3226 | Approved |
| 0.766 | Intermediate Similarity | NPD2798 | Approved |
| 0.7654 | Intermediate Similarity | NPD7473 | Discontinued |
| 0.7632 | Intermediate Similarity | NPD2533 | Approved |
| 0.7632 | Intermediate Similarity | NPD2534 | Approved |
| 0.7632 | Intermediate Similarity | NPD2532 | Approved |
| 0.7626 | Intermediate Similarity | NPD1481 | Phase 2 |
| 0.7619 | Intermediate Similarity | NPD7033 | Discontinued |
| 0.7619 | Intermediate Similarity | NPD4308 | Phase 3 |
| 0.7606 | Intermediate Similarity | NPD1530 | Clinical (unspecified phase) |
| 0.7589 | Intermediate Similarity | NPD3266 | Approved |
| 0.7589 | Intermediate Similarity | NPD3267 | Approved |
| 0.7576 | Intermediate Similarity | NPD7993 | Clinical (unspecified phase) |
| 0.7576 | Intermediate Similarity | NPD5953 | Discontinued |
| 0.7554 | Intermediate Similarity | NPD1535 | Discovery |
| 0.7552 | Intermediate Similarity | NPD4908 | Phase 1 |
| 0.7534 | Intermediate Similarity | NPD447 | Suspended |
| 0.7533 | Intermediate Similarity | NPD2654 | Approved |
| 0.7532 | Intermediate Similarity | NPD4288 | Approved |
| 0.7531 | Intermediate Similarity | NPD3926 | Phase 2 |
| 0.753 | Intermediate Similarity | NPD6559 | Discontinued |
| 0.7518 | Intermediate Similarity | NPD9545 | Approved |
| 0.7517 | Intermediate Similarity | NPD2346 | Discontinued |
| 0.75 | Intermediate Similarity | NPD1608 | Approved |
| 0.75 | Intermediate Similarity | NPD9493 | Approved |
| 0.7483 | Intermediate Similarity | NPD4628 | Phase 3 |
| 0.7466 | Intermediate Similarity | NPD1613 | Approved |
| 0.7466 | Intermediate Similarity | NPD1612 | Clinical (unspecified phase) |
| 0.7451 | Intermediate Similarity | NPD7390 | Discontinued |
| 0.7448 | Intermediate Similarity | NPD4907 | Clinical (unspecified phase) |
| 0.7448 | Intermediate Similarity | NPD411 | Approved |
| 0.7429 | Intermediate Similarity | NPD1201 | Approved |
| 0.7415 | Intermediate Similarity | NPD1899 | Clinical (unspecified phase) |
| 0.7415 | Intermediate Similarity | NPD1933 | Approved |
| 0.7415 | Intermediate Similarity | NPD230 | Phase 1 |
| 0.7413 | Intermediate Similarity | NPD1019 | Discontinued |
| 0.7407 | Intermediate Similarity | NPD1241 | Discontinued |
| 0.74 | Intermediate Similarity | NPD1471 | Phase 3 |
| 0.7394 | Intermediate Similarity | NPD3751 | Discontinued |
| 0.7371 | Intermediate Similarity | NPD8090 | Clinical (unspecified phase) |
| 0.7368 | Intermediate Similarity | NPD6398 | Clinical (unspecified phase) |
| 0.7356 | Intermediate Similarity | NPD4360 | Phase 2 |
| 0.7356 | Intermediate Similarity | NPD4363 | Phase 3 |
| 0.7349 | Intermediate Similarity | NPD5844 | Phase 1 |
| 0.7347 | Intermediate Similarity | NPD4307 | Phase 2 |
| 0.7343 | Intermediate Similarity | NPD1470 | Approved |
| 0.7329 | Intermediate Similarity | NPD6798 | Discontinued |
| 0.7329 | Intermediate Similarity | NPD3764 | Approved |
| 0.732 | Intermediate Similarity | NPD2354 | Approved |
| 0.731 | Intermediate Similarity | NPD1008 | Clinical (unspecified phase) |
| 0.7286 | Intermediate Similarity | NPD17 | Approved |
| 0.7279 | Intermediate Similarity | NPD6233 | Phase 2 |
| 0.7273 | Intermediate Similarity | NPD7784 | Clinical (unspecified phase) |
| 0.7256 | Intermediate Similarity | NPD3787 | Discontinued |
| 0.7254 | Intermediate Similarity | NPD3972 | Approved |
| 0.7233 | Intermediate Similarity | NPD7615 | Clinical (unspecified phase) |
| 0.7225 | Intermediate Similarity | NPD4287 | Approved |
| 0.7222 | Intermediate Similarity | NPD1164 | Approved |
| 0.7216 | Intermediate Similarity | NPD4361 | Phase 2 |
| 0.7216 | Intermediate Similarity | NPD4362 | Clinical (unspecified phase) |
| 0.7205 | Intermediate Similarity | NPD2296 | Approved |
| 0.7202 | Intermediate Similarity | NPD1729 | Discontinued |
| 0.7197 | Intermediate Similarity | NPD6980 | Clinical (unspecified phase) |
| 0.719 | Intermediate Similarity | NPD1652 | Phase 2 |
| 0.7181 | Intermediate Similarity | NPD5123 | Clinical (unspecified phase) |
| 0.7181 | Intermediate Similarity | NPD5124 | Phase 1 |
| 0.7179 | Intermediate Similarity | NPD7004 | Clinical (unspecified phase) |
| 0.7171 | Intermediate Similarity | NPD2355 | Clinical (unspecified phase) |
| 0.7171 | Intermediate Similarity | NPD2353 | Approved |
| 0.7169 | Intermediate Similarity | NPD2403 | Approved |
| 0.7164 | Intermediate Similarity | NPD9266 | Approved |
| 0.7164 | Intermediate Similarity | NPD74 | Approved |
| 0.7143 | Intermediate Similarity | NPD4625 | Phase 3 |
| 0.7143 | Intermediate Similarity | NPD1894 | Discontinued |
| 0.709 | Intermediate Similarity | NPD9263 | Approved |
| 0.709 | Intermediate Similarity | NPD9267 | Approved |
| 0.709 | Intermediate Similarity | NPD9265 | Clinical (unspecified phase) |
| 0.709 | Intermediate Similarity | NPD9264 | Approved |
| 0.7083 | Intermediate Similarity | NPD4749 | Approved |
| 0.7081 | Intermediate Similarity | NPD6844 | Discontinued |
| 0.7067 | Intermediate Similarity | NPD4340 | Discontinued |
| 0.7067 | Intermediate Similarity | NPD6355 | Discontinued |
| 0.7051 | Intermediate Similarity | NPD7440 | Discontinued |
| 0.7048 | Intermediate Similarity | NPD5710 | Approved |
| 0.7048 | Intermediate Similarity | NPD5711 | Approved |
| 0.7045 | Intermediate Similarity | NPD7879 | Clinical (unspecified phase) |
| 0.7025 | Intermediate Similarity | NPD5049 | Phase 3 |
| 0.7018 | Intermediate Similarity | NPD6104 | Discontinued |
| 0.7015 | Intermediate Similarity | NPD968 | Approved |
| 0.7014 | Intermediate Similarity | NPD9269 | Phase 2 |
| 0.7007 | Intermediate Similarity | NPD2861 | Phase 2 |
| 0.7006 | Intermediate Similarity | NPD6143 | Clinical (unspecified phase) |
| 0.7006 | Intermediate Similarity | NPD7184 | Clinical (unspecified phase) |
| 0.7 | Intermediate Similarity | NPD405 | Clinical (unspecified phase) |
| 0.6993 | Remote Similarity | NPD5404 | Approved |
| 0.6993 | Remote Similarity | NPD5406 | Approved |
| 0.6993 | Remote Similarity | NPD5408 | Approved |
| 0.6993 | Remote Similarity | NPD5405 | Approved |
| 0.6987 | Remote Similarity | NPD3887 | Approved |
| 0.6987 | Remote Similarity | NPD6190 | Approved |
| 0.6982 | Remote Similarity | NPD5940 | Clinical (unspecified phase) |
| 0.6972 | Remote Similarity | NPD9268 | Approved |
| 0.6957 | Remote Similarity | NPD6585 | Discontinued |
| 0.6948 | Remote Similarity | NPD2343 | Clinical (unspecified phase) |
| 0.6946 | Remote Similarity | NPD7229 | Phase 3 |
| 0.6937 | Remote Similarity | NPD1653 | Approved |
| 0.6933 | Remote Similarity | NPD4062 | Phase 3 |
| 0.6919 | Remote Similarity | NPD7584 | Approved |
| 0.6918 | Remote Similarity | NPD1876 | Approved |
| 0.6914 | Remote Similarity | NPD5889 | Approved |
| 0.6914 | Remote Similarity | NPD5890 | Approved |
| 0.6914 | Remote Similarity | NPD8434 | Phase 2 |
 Bioactivity similarity: Similar Natural Products in NPASS
Bioactivity similarity: Similar Natural Products in NPASSBioactivity similarity was calculated based on bioactivity descriptors of compounds. The bioactivity descriptors were calculated by a recently developed AI algorithm Chemical Checker (CC) [Nature Biotechnology, 38:1087–1096, 2020; Nature Communications, 12:3932, 2021], which evaluated bioactivity similarities at five levels:
☉ A: chemistry similarity;
☉ B: biological targets similarity;
☉ C: networks similarity;
☉ D: cell-based bioactivity similarity;
☉ E: similarity based on clinical data.
Those 5 categories of CC bioactivity descriptors were calculated and then subjected to manifold projection using UMAP algorithm, to project all NPs on a 2-Dimensional space. The current NP was highlighted with a small circle in the 2-D map. Below figures: left-to-right, A-to-E.